1 Simulation example Q.6
In this simulation example, the highlighted data analysis pitfall is sampling variability. Specifically, it shows the influence of sampling variability on the results of a data analysis.
1.1 Data-generating process
The data-generating process is the specific process that generates the observed data. In reality, this process is unknown and understanding it is, in a general sense, the ultimate aim of science. Here, we assume this process is known and describe it via a directed acyclic graph (DAG) in Fig. 1.1. In this DAG,1 indoor air temperature \((\text{T})\) and biological sex \((\text{S})\) influence thermal sensation vote \((\text{V})\) both directly and indirectly, passing through clothing insulation \((\text{C})\).
Code
dag_coords <-
data.frame(name = c('T', 'C', 'S', 'V'),
x = c(1, 2, 3, 2),
y = c(1.2, 1.2, 1.2, 1))
DAG.06 <-
dagify(C ~ S + T,
V ~ S + C + T,
coords = dag_coords)
ggplot(data = DAG.06, aes(x = x, y = y, xend = xend, yend = yend)) +
geom_dag_text(colour='black', size = 8, parse = TRUE,
family = c('mono'),
label = c(expression(bold(C)), expression(bold(S)), expression(bold(T)), expression(bold(V)))) +
geom_dag_edges(arrow_directed = grid::arrow(length = grid::unit(8, 'pt'), type = 'open'),
family = c('mono'),
fontface = c('bold')) +
coord_cartesian(xlim = c(1, 3), ylim = c(1, 1.2)) +
theme_dag()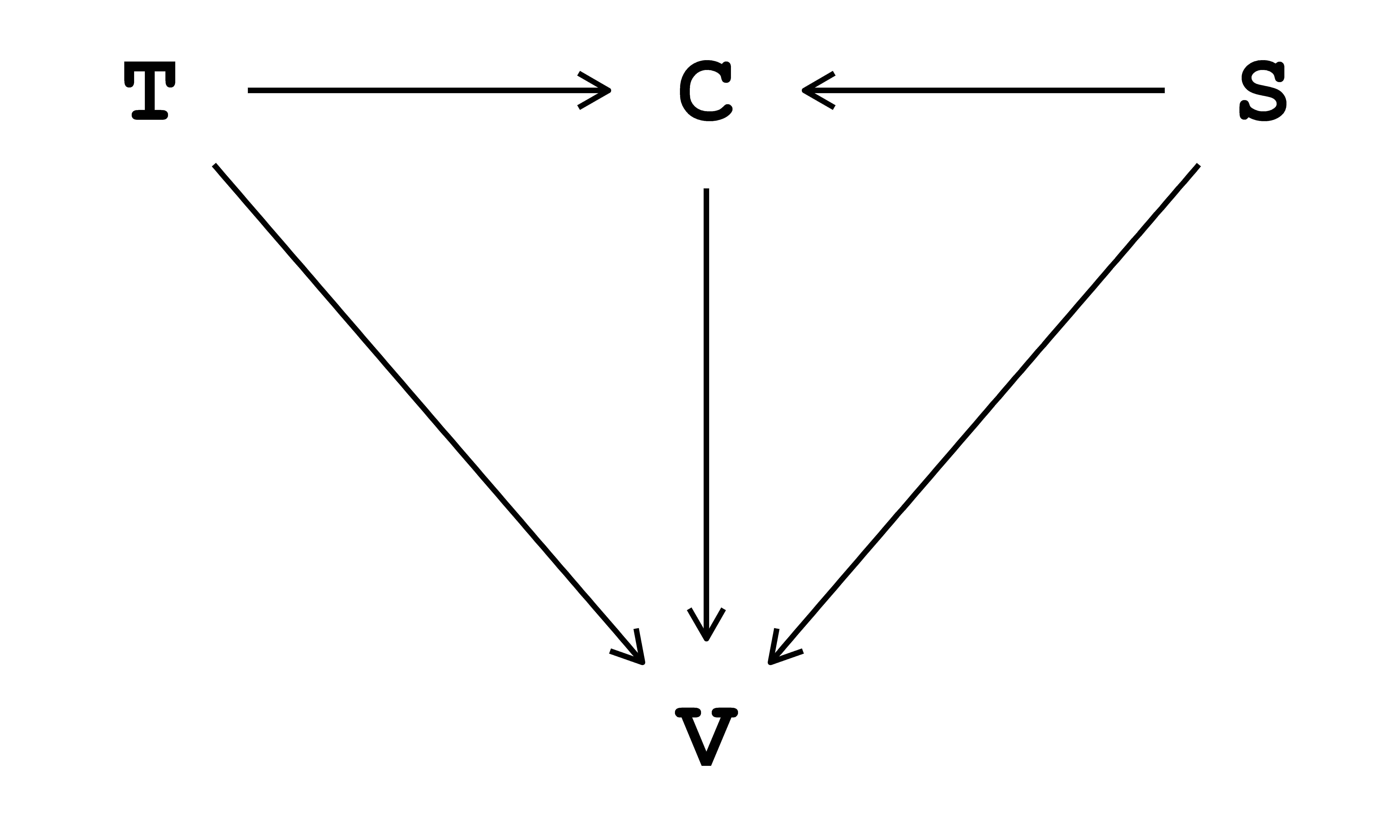
The DAG in Fig. 1.1 can be written as:
- \(C \sim f_{C}(S, T)\), read as ‘clothing insulation \((\text{C})\) is some function of biological sex \((\text{S})\) and indoor air temperature \((\text{T})\)’.
- \(V \sim f_{V}(S, T, C)\), read as ‘thermal sensation vote \((\text{V})\) is some function of biological sex \((\text{S})\), indoor air temperature \((\text{T})\) and clothing insulation \((\text{C})\)’.
1.2 Synthetic data set
In a DAG, an omitted arrow between two variables signifies the absence of direct influence of one variable on another. In contrast, a present arrow does not provide any information about the magnitude of the effect or its functional form. As such, all relationships are qualitative. However, to generate synthetic data, we need to quantitatively define the relationship between the variables. These relationships are defined within the custom function data.sim_Q06.ord(). This function takes as input the sample size N and generates synthetic data according to the DAG in Fig. 1.1.
data.sim_Q06.ord <- function(N) {
#Simulate biological sex
S <- factor(sample(1:2, size = N, replace = TRUE)) #S = 1 'male'; S = 2 'female'
#Simulate indoor air temperature
T <- rnorm(N, mean = 23 , sd = 2)
#Simulate clothing insulation
C <- rlnormTrunc(N, meanlog = ifelse(S=='1', 0.25 + 0.00 -0.04*T, 0.25 + 0.03 -0.04*T), sdlog = 0.15, min = 0.3 , max = 2)
#Simulate thermal sensation vote
V <- sim_TSV.ord(n = N, sex = S, temp = T, clo = C, b_int = 0, b_sex = c(0,-0.16), b_temp = 0.37, b_clo = 0.88)
return(data.table(S, T, C, V)) #return a data.table with the simulated values
}Specifically:
Biological sex \((\text{S})\) is randomly sampled with replacement from a vector of factors
1(i.e., male) and2(i.e., female).Indoor air temperature \((\text{T})\) is randomly sampled from a normal distribution. The mean is set to 23°C (i.e.,
mean = 23) and the standard deviation to 2°C (i.e.,sd = 2).Clothing insulation \((\text{C})\) is randomly sampled from a truncated lognormal distribution. The mean is on the log scale (i.e.,
meanlog) and is a linear of function of biological sex \((\text{S})\) and indoor air temperature \((\text{T})\): for male (i.e.,S = 1) ismeanlog = 0.25 - 0.04*Twhile for female (i.e.,S = 2) ismeanlog = 0.25 + 0.03 - 0.04*T. The standard deviation is on the log scale and is a constant, fixed at 0.15 (i.e.,sdlog = 0.15). The parametersminandmaxare the assumed lower and upper bounds for clothing, fixed to 0.3 clo (i.e.,min = 0.3) and 2 clo (i.e.,max = 2), respectively.Thermal sensation votes \((\text{V})\) are generated from the custom function
sim_TSV.ord(), which is a function of biological sex (sex), indoor air temperature (temp), and clothing insulation (clo). The termb_intis the coefficient for the intercept (set to0),b_sexis the coefficient for biological sex (set to0for males and-0.16for females),b_tempis the coefficient for indoor air temperature (set to0.37), andb_clois the coefficient for clothing insulation (set to0.88). More information on the custom functionsim_TSV.ord()can be found in Appendix A.1.
From this data generation mechanism, we simulated the target population, which consists of one million observations.
set.seed(2023) #Set random number for reproducibility
#Simulate the population
Q06_population <- data.sim_Q06.ord(N = 1e6)
#Transform V to an ordered factor
Q06_population[, V := factor(V, levels = c('1','2','3','4','5','6','7'), ordered = TRUE)]
#View the data frame
Q06_population S T C V
<fctr> <num> <num> <ord>
1: 1 19.35834 0.5173312 3
2: 2 20.17534 0.6351932 3
3: 1 17.25539 0.6350222 4
4: 1 22.32519 0.5410619 4
5: 2 23.51410 0.5319473 6
---
999996: 1 23.16175 0.5811199 7
999997: 1 20.21810 0.4694916 4
999998: 2 21.43980 0.5467828 3
999999: 1 21.73996 0.4982545 4
1000000: 1 20.66438 0.6083867 3From this population, we obtained three thousand data sets with three different sample sizes using simple random sampling. Specifically, a thousand data sets with sample sizes of 30, 300 and 900.
#Sample one data set of 30 observations
set.seed(random.seed[1])
sample.random_3e1 <-
Q06_population |>
slice_sample(n = 3e1) #take a simple random sample of size 30
#View the data frame
sample.random_3e1 S T C V
<fctr> <num> <num> <ord>
1: 2 21.09664 0.5414387 2
2: 1 21.71421 0.5381658 4
3: 2 23.74820 0.5389922 6
4: 2 20.05361 0.5974996 4
5: 1 20.72877 0.5577154 3
6: 2 24.28257 0.6013693 5
7: 1 24.92904 0.4196810 4
8: 1 21.80931 0.6892414 4
9: 1 21.82005 0.4485176 3
10: 1 26.24905 0.4085024 6
11: 2 27.07587 0.5773195 5
12: 2 23.37962 0.4209909 4
13: 2 24.04006 0.5938105 4
14: 2 24.71866 0.5924425 4
15: 1 26.38131 0.3865448 2
16: 2 20.23951 0.5525658 4
17: 1 25.08493 0.4105008 4
18: 2 21.63454 0.4554378 4
19: 2 22.27104 0.5191736 4
20: 2 23.71114 0.6124570 2
21: 2 28.60947 0.4721329 5
22: 2 22.36287 0.5316676 4
23: 1 20.99654 0.6277547 4
24: 1 23.20989 0.5144766 3
25: 1 21.83795 0.5868481 4
26: 1 22.51528 0.6097533 4
27: 1 23.08223 0.4722961 4
28: 2 23.65405 0.4397346 3
29: 1 26.82511 0.4086332 4
30: 1 21.13123 0.5078074 4
S T C V#Sample one data set of 300 observations
set.seed(random.seed[1])
sample.random_3e2 <-
Q06_population |>
slice_sample(n = 3e2) #take a simple random sample of size 300
#View the data frame
sample.random_3e2 S T C V
<fctr> <num> <num> <ord>
1: 2 21.09664 0.5414387 2
2: 1 21.71421 0.5381658 4
3: 2 23.74820 0.5389922 6
4: 2 20.05361 0.5974996 4
5: 1 20.72877 0.5577154 3
---
296: 2 22.08967 0.7875628 4
297: 1 23.89738 0.3811996 4
298: 2 21.42748 0.5020544 3
299: 1 21.06568 0.4899180 4
300: 2 19.44767 0.5651442 1#Sample one data set of 900 observations
set.seed(random.seed[1])
sample.random_9e2 <-
Q06_population |>
slice_sample(n = 9e2) #take a simple random sample of size 900
#View the data frame
sample.random_9e2 S T C V
<fctr> <num> <num> <ord>
1: 2 21.09664 0.5414387 2
2: 1 21.71421 0.5381658 4
3: 2 23.74820 0.5389922 6
4: 2 20.05361 0.5974996 4
5: 1 20.72877 0.5577154 3
---
896: 1 22.01716 0.6665051 4
897: 2 25.58033 0.4767277 4
898: 1 21.13034 0.6622662 4
899: 1 20.12777 0.6227197 5
900: 1 21.80400 0.4828775 4A summary of descriptive statistics for all variables in the data sets is given below; of the three thousand data sets, only one for each of the three sample sizes (i.e., 30, 300 and 900) is presented. Detailed information can be found in Q.6 Summary.
summary(sample.random_3e1) S T C V
1:15 Min. :20.05 Min. :0.3865 1: 0
2:15 1st Qu.:21.74 1st Qu.:0.4502 2: 3
Median :23.15 Median :0.5349 3: 4
Mean :23.31 Mean :0.5211 4:18
3rd Qu.:24.61 3rd Qu.:0.5910 5: 3
Max. :28.61 Max. :0.6892 6: 2
7: 0 summary(sample.random_3e2) S T C V
1:156 Min. :17.45 Min. :0.3393 1: 7
2:144 1st Qu.:21.74 1st Qu.:0.4571 2: 15
Median :23.07 Median :0.5126 3: 39
Mean :23.03 Mean :0.5195 4:157
3rd Qu.:24.31 3rd Qu.:0.5792 5: 54
Max. :29.57 Max. :0.7876 6: 18
7: 10 summary(sample.random_9e2) S T C V
1:447 Min. :15.49 Min. :0.3091 1: 14
2:453 1st Qu.:21.70 1st Qu.:0.4613 2: 29
Median :23.05 Median :0.5175 3:160
Mean :23.02 Mean :0.5242 4:437
3rd Qu.:24.36 3rd Qu.:0.5826 5:185
Max. :29.57 Max. :0.8901 6: 49
7: 26 1.3 Data analysis
In a scientific context, data analysis should be performed to model the phenomenon under scrutiny and not fitting curves to data. The data are a means of obtaining information about that phenomenon to answer the specific question of interest. In this example, we assumed that the objective of the data analysis is to answer the following inferential question: ‘Is there a direct association between biological sex and thermal sensation vote?’.
1.3.1 Scientific model
The first step to address a scientific question is to define the scientific model. A scientific model is essentially a representation of a specific phenomenon that is being studied. To achieve this, we defined a DAG. A DAG generally consists of letter-and-arrow pictures summarising the researcher’s scientific knowledge about the phenomenon under study. Since we know the data-generating process (we defined it), the scientific model equates to the data-generating process outlined in Fig. 1.1. In reality, however, it would not.2
1.3.2 Statistical model
A statistical model is a mathematical representation of the scientific model —which in turn represents the data-generating process— that aims to answer the particular question of interest. To address the question, the synthetic data sets are analysed using ordinal regression within a frequentist framework.3
To estimate the effect of interest, we need to select the variables to adjust for: the adjustment set. Assuming that the structure of the scientific model is correct, we can use the adjustmentSets() function to identify the adjustment set that (asymptotically) allow the unbiased estimation of the effect of interest.
adjustmentSets(DAG.06,
exposure = 'S',
outcome = 'V',
type = 'minimal',
effect = 'direct',
max.results = Inf){ C, T }The resulting adjustment set include both clothing \((\text{C})\) and indoor air temperature \((\text{T})\). The adjustment set is only one for this DAG, but there can be several adjustment sets for a more complex DAG.
The statistical model can then be written as:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Multinomial}(1, \boldsymbol\pi_{i})\\ \operatorname{logit}(\gamma_{ik}) &= \tau_{k} -\eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{S} (\mathrm{S}_i) + \beta_{T} \mathrm{T}_{i} + \beta_{C} \mathrm{C}_{i}\\ \end{aligned} \tag{1.1}\]
In Eq. 1.1, the conditional distribution of the response variable \(\mathrm{Y}_{i}\) for the \(i^{th}\) observation is assumed to follow an ordered multinomial distribution with probability vector \(\boldsymbol\pi_{i} = \{\pi_{i1},\ ...,\ \pi_{ik}\}\), where \(\pi_{ik} = \mathrm{Pr}(Y_{i}=k)\). The cumulative probability for \(\pi_{ik}\) is \(\gamma_{ik} = \mathrm{Pr}(Y_{i} \le k)\), hence \(\gamma_{ik} = \pi_{i1}\ + ... +\ \pi_{ik}.\) The term \(\{\tau_{k}\}\) are the threshold parameters, with \(k \in \{1,\ ...,\ K\}\) and \(\eta_{i}\) is the linear predictor, where \(\alpha_{0}\) is the intercept term4 and \(\beta_{S}\), \(\beta_{T}\), \(\beta_{C}\) are the regression coefficients for biological sex \((\text{S})\), indoor air temperature \((\text{T})\), and clothing insulation \((\text{C})\), respectively.
A summary description of the simulation example is provided in Table 1.1.
| Pitfall | Forgetting the influence of sampling variability on the results and interpretation of a data analysis. |
| Type of analysis | Inferential. |
| Research question | Is there a direct association between biological sex and TSV? |
| Framework | Frequentist. |
| Assumptions | Random sample (simple random sampling): everyone in the population has an equal chance of being selected into the sample. |
| Independence of observations: each observation represents independent bits of information. | |
| No confounding: the DAG includes all shared causes among the variables. | |
| No model error: perfect functional form specification. | |
| No measurement error: all variables are measured perfectly. | |
| Variables | Indoor air temperature (T): continuous variable [unit: °C] |
| Thermal resistance of clothing (C): continuous variable [unit: clo] | |
| Sex (S): categorical variable [‘1’ male; ‘2’ female] | |
| TSV (V): ordinal variable [‘1’ cold; ‘2’ cool; ‘3’ slightly cool; ‘4’ neutral; ‘5’ slightly warm; ‘6’ warm; ‘7’ hot] |
1.3.3 Results
The results of the fitted statistical model (defined above) are presented here. The for-loop shown in the code below performs the following operations. First samples (using simple random sample) a data set of 30 observations from the target population. Subsequently, perform ordinal regression and store the estimated coefficient for sex and its 95% confidence interval in the data frame coefs_3e1. This operation is repeated a thousand times, resulting in the data frame coefs_3e1 containing the estimates (point estimate and confidence interval) of a thousand random samples of size 30.
#Create an empty data frame
empty.df <- data.frame(matrix(NA, nrow = n_sims, ncol = 6))
#Rename the data frame columns
colnames(empty.df) <- c('sim.id', 'estimate', 'CI_2.5', 'CI_97.5',
'sample.size', 'coverage')
#Sample a thousand data sets of 30 observations and perform ordinal regression
coefs_3e1 <- empty.df #assign the empty data frame
for (i in 1:n_sims){
set.seed(random.seed[i]) #set unique seed for each simulation
#Sample data set from population
sample.random <-
Q06_population |>
slice_sample(n = 3e1) #take a simple random sample of size 30
#Fit model
fit <- clm(formula = V ~ 1 + S + T + C,
data = sample.random,
link = 'logit',
threshold = 'flexible')
#Compile matrix
coefs_3e1[i, 1] <- i #simulation ID
coefs_3e1[i, 2] <- coef(fit)['S2'] #point estimate
coefs_3e1[i, 3:4] <- confint(fit, level = 0.95, type = 'Wald')['S2', ] #confidence interval (Wald)
coefs_3e1[i, 5] <- '3e1' #sample size
}
#View the first six rows of the data frame
head(coefs_3e1, 6) sim.id estimate CI_2.5 CI_97.5 sample.size coverage
1 1 0.4111986 -1.0894662 1.9118634 3e1 NA
2 2 0.6224043 -0.9730538 2.2178624 3e1 NA
3 3 -0.4799724 -1.9708800 1.0109352 3e1 NA
4 4 0.4042922 -1.1084313 1.9170156 3e1 NA
5 5 -1.9331813 -3.6423774 -0.2239852 3e1 NA
6 6 0.4240608 -1.0861795 1.9343011 3e1 NAThe same procedure is repeated for the data sets of sample sizes of 300 and 900, and the results are stored in coefs_3e2 and coefs_9e2, respectively.
Code
#Sample a thousand data sets of 300 observations and perform ordinal regression
coefs_3e2 <- empty.df #assign the empty data frame
for (i in 1:n_sims){
set.seed(random.seed[i]) #set unique seed for each simulation
#Sample data set from population
sample.random <-
Q06_population |>
slice_sample(n = 3e2) #take a simple random sample of size 300
#Fit model
fit <- clm(formula = V ~ 1 + S + T + C,
data = sample.random,
link = 'logit',
threshold = 'flexible')
#Compile matrix
coefs_3e2[i, 1] <- i #simulation ID
coefs_3e2[i, 2] <- coef(fit)['S2'] #point estimate
coefs_3e2[i, 3:4] <- confint(fit, level = 0.95, type = 'Wald')['S2', ] #confidence interval (Wald)
coefs_3e2[i, 5] <- '3e2' #sample size
}
#Sample a thousand data sets of 900 observations and perform ordinal regression
coefs_9e2 <- empty.df #assign the empty data frame
for (i in 1:n_sims){
set.seed(random.seed[i]) #set unique seed for each simulation
#Sample data set from population
sample.random <-
Q06_population |>
slice_sample(n = 9e2) #take a simple random sample of size 900
#Fit model
fit <- clm(formula = V ~ 1 + S + T + C,
data = sample.random,
link = 'logit',
threshold = 'flexible')
#Compile matrix
coefs_9e2[i, 1] <- i #simulation ID
coefs_9e2[i, 2] <- coef(fit)['S2'] #point estimate
coefs_9e2[i, 3:4] <- confint(fit, level = 0.95, type = 'Wald')['S2', ] #confidence interval (Wald)
coefs_9e2[i, 5] <- '9e2' #sample size
}The three data frames (coefs_3e1, coefs_3e2, and coefs_9e2) are merged (i.e., coefs), and the coverage is calculated. The coverage is set to 1 if the confidence interval overlaps the data-generating parameter for sex (i.e., b_sex = -0.16) and 0 otherwise.
b_sex <- c(0,-0.16) # Male, Female
#Combine the three data frames
coefs <- data.table(rbind(coefs_3e1, coefs_3e2, coefs_9e2))
#Calculate coverage
coefs[, coverage := as.character(coverage)][, coverage := '1'][(b_sex[2] < CI_2.5) | (b_sex[2] > CI_97.5), coverage := '0']
#View the data frame
coefs sim.id estimate CI_2.5 CI_97.5 sample.size coverage
<int> <num> <num> <num> <char> <char>
1: 1 0.4111986 -1.0894662 1.91186336 3e1 1
2: 2 0.6224043 -0.9730538 2.21786242 3e1 1
3: 3 -0.4799724 -1.9708800 1.01093518 3e1 1
4: 4 0.4042922 -1.1084313 1.91701563 3e1 1
5: 5 -1.9331813 -3.6423774 -0.22398515 3e1 0
---
2996: 996 -0.1039069 -0.3491340 0.14132018 9e2 1
2997: 997 -0.3088756 -0.5543428 -0.06340832 9e2 1
2998: 998 -0.5290883 -0.7817956 -0.27638097 9e2 0
2999: 999 0.0849292 -0.1634437 0.33330208 9e2 1
3000: 1000 -0.1757759 -0.4226830 0.07113115 9e2 1The results are then plotted in Fig. 1.2.
Code
ggplot(data = subset(coefs, sim.id <= 1e2), aes(x = sim.id, y = estimate, ymin = CI_2.5, ymax = CI_97.5, colour = coverage)) +
geom_hline(yintercept = b_sex[2], alpha = 0.8, linetype = 'dashed', colour = 'blue') +
geom_linerange() +
geom_point(size = 1) +
scale_colour_manual('', values = c('black', '#FF4040'),
breaks=c('1','0')) +
scale_y_continuous('Estimate (logit scale)') +
scale_x_continuous('Simulation ID') +
facet_wrap(.~factor(sample.size,
levels=c('3e1', '3e2', '9e2'),
labels = c('Sample size = 30', 'Sample size = 300', 'Sample size = 900')),
nrow = 3,
scales = 'fixed') +
theme(legend.position = 'none',
axis.title.x = element_text(margin = unit(c(2, 0, 0, 0), "mm")),
axis.title.y = element_text(margin = unit(c(0, 2, 0, 0), "mm"))
)
Fig. 1.2 shows the first 100 estimates (point estimate and confidence interval) of the coefficient for sex for the three different sample sizes. The inferential uncertainty shrinks (i.e., narrow confidence intervals) when the sample size increases from 30 to 900 observations. However, it can be seen that, independently of the sample size, some confidence intervals (the one in red) do not overlap the data-generating parameter (the dashed blue line). If all the thousand simulations are considered, the frequency of the coverage of the calculated confidence intervals (i.e., how many times the confidence intervals overlap the data-generating parameter) is 92.4%, 95.7% and 95.8% for the 30, 300 and 900 observations, respectively. This result is expected because it is a property that a 95% confidence interval has. However, once a specific confidence interval has been calculated (in our case, the result of one simulation), it is impossible to make any conclusions about the probability of it containing the data-generating parameter. In the frequentist approach probability refers to frequency. The data-generating parameter is a fixed constant, and a calculated confidence interval is also fixed. Therefore, a given confidence interval either includes the parameter or does not, with no frequency involved. Confidence intervals only quantify the uncertainty due to random error (i.e., sample variability), not systematic error (i.e., bias). According to Gelman et al. (pages 56 of Ref. (Gelman et al., 2020)), the ways to account for sources of errors that are not in the statistical model are: (i) improving data collection, (ii) expanding the model, and (iii) increasing the stated uncertainty. Concerning the last point, some authors (see Ref. (Lash et al., 2009) and chapter 19 of Ref. (Rothman et al., 2008)) recommend the application of quantitative bias analysis to produce intervals around the effect estimate, taking into account random and systematic sources of uncertainty.
When conducting a study, it is important to recognise that (generally) only uncertain conclusions can be drawn from data, even if statistical significance is achieved. In thermal comfort, claims on data tend to be stated deterministically by placing the results into the ‘significant’ or ‘nonsignificant’ bins. However, this practice, called dichotomania (Greenland, 2017), is harmful because it overlooks the inherent variation in the human response (but more generally in the topic under study) as well as the uncertainty involved in statistical inference. Instead, focusing on uncertainty quantification is likely to result in a reduction of overly confident assertions that, upon further examination, may lack support from the data. Practical examples of this suggestion can be found in Vasishth and Gelman (Vasishth and Gelman, 2021).
Session info
Version information about R, the OS and attached or loaded packages.
R version 4.2.3 (2023-03-15 ucrt)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 22631)
Matrix products: default
locale:
[1] LC_COLLATE=English_United Kingdom.utf8
[2] LC_CTYPE=English_United Kingdom.utf8
[3] LC_MONETARY=English_United Kingdom.utf8
[4] LC_NUMERIC=C
[5] LC_TIME=English_United Kingdom.utf8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] EnvStats_2.8.1 ordinal_2023.12-4 lubridate_1.9.3 forcats_1.0.0
[5] stringr_1.5.1 dplyr_1.1.4 purrr_1.0.2 readr_2.1.5
[9] tidyr_1.3.1 tibble_3.2.1 tidyverse_2.0.0 gt_0.11.0
[13] data.table_1.15.4 ggplot2_3.5.1 dagitty_0.3-4 ggdag_0.2.12
loaded via a namespace (and not attached):
[1] ggrepel_0.9.5 Rcpp_1.0.12 lattice_0.20-45
[4] digest_0.6.36 utf8_1.2.4 V8_4.4.2
[7] ggforce_0.4.2 R6_2.5.1 evaluate_0.24.0
[10] pillar_1.9.0 rlang_1.1.4 curl_5.2.1
[13] rstudioapi_0.16.0 Matrix_1.6-5 rmarkdown_2.27
[16] labeling_0.4.3 htmlwidgets_1.6.4 igraph_2.0.3
[19] polyclip_1.10-6 munsell_0.5.1 compiler_4.2.3
[22] numDeriv_2016.8-1.1 xfun_0.46 pkgconfig_2.0.3
[25] htmltools_0.5.8.1 tidyselect_1.2.1 gridExtra_2.3
[28] graphlayouts_1.1.1 viridisLite_0.4.2 fansi_1.0.6
[31] ucminf_1.2.2 tzdb_0.4.0 withr_3.0.0
[34] MASS_7.3-58.2 grid_4.2.3 nlme_3.1-162
[37] jsonlite_1.8.8 gtable_0.3.5 lifecycle_1.0.4
[40] magrittr_2.0.3 scales_1.3.0 cachem_1.1.0
[43] cli_3.6.2 stringi_1.8.4 farver_2.1.2
[46] viridis_0.6.5 xml2_1.3.6 generics_0.1.3
[49] vctrs_0.6.5 boot_1.3-28.1 tools_4.2.3
[52] glue_1.7.0 tweenr_2.0.3 hms_1.1.3
[55] ggraph_2.2.1 fastmap_1.2.0 yaml_2.3.9
[58] timechange_0.3.0 colorspace_2.1-0 tidygraph_1.3.1
[61] memoise_2.0.1 knitr_1.48 sass_0.4.9 We have selected only a few variables of interest to limit the complexity of the DAG and shift the focus to statistical data analysis.↩︎
The scientific model is generally an approximation (a best guess) of the unknown data-generating process. As such, it would have some inaccuracies/errors, which will inevitably affect the data analysis.↩︎
In all simulation examples, we relied on the regression framework (ordinal or beta regression) within the frequentist or Bayesian approach. However, other approaches are available and could be used depending on the question and the data. It is the researcher’s responsibility to justify the choice of the statistical framework (and any decisions made during the modelling process) to answer the specific question of interest.↩︎
Here, the intercept is omitted (i.e., set to zero). Omitting the intercept term allows the full set of thresholds \(\tau_{1},\ ...,\ \tau_{k}\) to be identified.↩︎