5 Simulation example Q.10
In this simulation example, the highlighted data analysis pitfall is random sampling variability. Specifically, it shows the influence of random sampling variability on the results of a data analysis.
5.1 Data-generating process
The data-generating process is the specific process that generates the observed data. In reality, this process is unknown and understanding it is, in a general sense, the ultimate aim of science. Here, we assume this process is known and describe it via a directed acyclic graph (DAG) in Fig. 5.1. In this DAG,1 indoor air temperature \((\text{T})\) and biological sex \((\text{S})\) influence thermal sensation vote \((\text{V})\) both directly and indirectly, passing through clothing insulation \((\text{C})\). The true indoor air temperature \(\text{T}\) cannot be observed, so it is circled as an unobserved node. However, we do get to observe \(\mathrm{T^*}\), which is a function of both the true air temperature \(\text{T}\) and some unobserved error \(\mathrm{e_{T}}\). The error itself is not directly observable but could be roughly approximated by considering the possible uncertainty in the measurement process.
Code
dag_coords <-
data.frame(name = c('T', 'C', 'S', 'V', 'Tobs', 'eT'),
x = c(1, 2, 3, 2, 1, 1.5),
y = c(1.2, 1.2, 1.2, 1, 1.3, 1.3))
DAG.10 <-
dagify(C ~ S,
C ~ T,
Tobs ~ T,
Tobs ~ eT,
V ~ S + C + T,
coords = dag_coords)
ggplot(data = DAG.10, aes(x = x, y = y, xend = xend, yend = yend)) +
geom_point(x = 1, y = 1.2,
shape = 1, size = 14,
stroke = 0.9, color = 'black') +
geom_point(x = 1.5, y = 1.3,
shape = 1, size = 14,
stroke = 0.9, color = 'black') +
geom_dag_text(colour='black', size = 8, parse = TRUE,
family = c('mono'),
label = c(expression(bold(C)), expression(bold(S)), expression(bold(T)),
expression(bold(T^scriptstyle('*'))), expression(bold(V)), expression(bold(e[T])))) +
geom_dag_edges(arrow_directed = grid::arrow(length = grid::unit(8, 'pt'), type = 'open'),
family = c('mono'),
fontface = c('bold')) +
coord_cartesian(xlim = c(1, 3), ylim = c(1, 1.3)) +
theme_dag()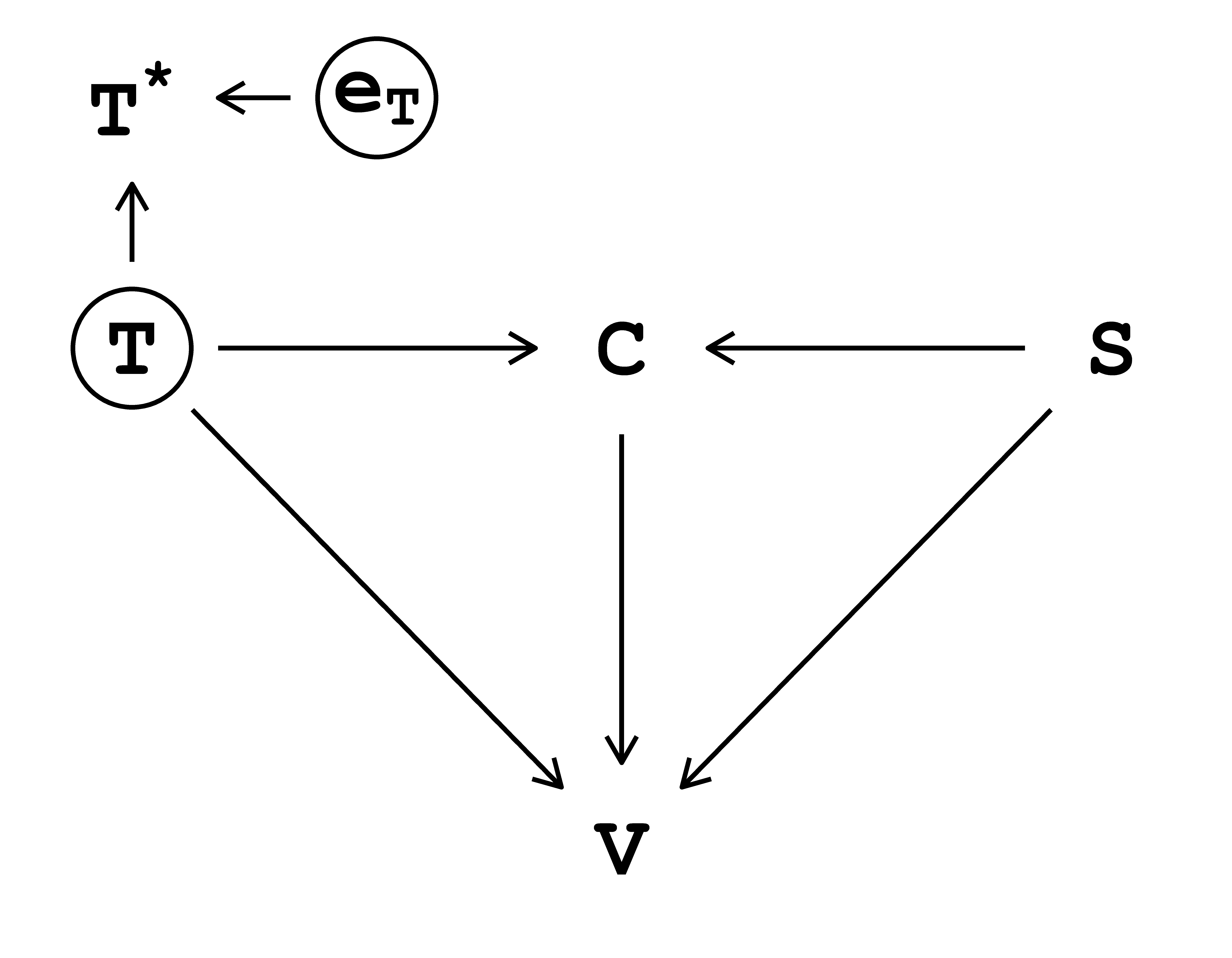
The DAG in Fig. 5.1 can be written as:
- \(T^* \sim f_{T^*}(T, e_{T})\), read as ‘The observable indoor air temperature \((\text{T})\) is some function of the true but unobservable air temperature \(\text{T}\) and some unobserved error \(\mathrm{e_{T}}\)’.
- \(C \sim f_{C}(S, T)\), read as ‘clothing insulation \((\text{C})\) is some function of biological sex \((\text{S})\) and indoor air temperature \((\text{T})\)’.
- \(V \sim f_{V}(S, T, C)\), read as ‘thermal sensation vote \((\text{V})\) is some function of biological sex \((\text{S})\), indoor air temperature \((\text{T})\) and clothing insulation \((\text{C})\)’.
5.2 Synthetic data set
In a DAG, an omitted arrow between two variables signifies the absence of direct influence of one variable on another. In contrast, a present arrow does not provide any information about the magnitude of the effect or its functional form. As such, all relationships are qualitative. However, to generate synthetic data, we need to quantitatively define the relationship between the variables. These relationships are defined within the custom function data.sim_Q10.beta(). This function takes as input the sample size N and generates synthetic data according to the DAG in Fig. 5.1.
data.sim_Q10.beta <- function(N) {
#Simulate biological sex
S <- as.factor(sample(1:2, size=N, replace=TRUE)) #S = 1 'male'; S = 2 'female'
#Simulate indoor air temperature
T <- rnorm(N, mean = 23 , sd = 2)
#Simulate clothing insulation
C <- rlnormTrunc(N, meanlog = ifelse(S=='1', 0.25 + 0.00 -0.04*T, 0.25 + 0.03 -0.04*T), sdlog = 0.15, min = 0.3 , max = 2) #0.15
#Simulate various measurement error
e_T.00 <- 0 #no measurement error
e_T.01 <- rnorm(N, mean = 0, sd = 0.10) #measurement error with 0 mean and 0.10 standard deviation
e_T.02 <- rnorm(N, mean = 0, sd = 0.20) #measurement error with 0 mean and 0.20 standard deviation
e_T.03 <- rnorm(N, mean = 0, sd = 0.30) #measurement error with 0 mean and 0.30 standard deviation
e_T.04 <- rnorm(N, mean = 0, sd = 0.40) #measurement error with 0 mean and 0.40 standard deviation
e_T.05 <- rnorm(N, mean = 0, sd = 0.50) #measurement error with 0 mean and 0.50 standard deviation
#Measurement error assumed during the analysis
sd_T <- rep(0.23, times = N)
#Simulate thermal sensation vote
V <- sim_TSV.beta(n=N, sex=S, temp=T, clo=C, b_int=-8.65, b_sex=c(0,-0.16), b_temp=0.37, b_clo=0.88, phi=17.9)
return(data.table(S, T, C, e_T.00, e_T.01, e_T.02, e_T.03, e_T.04, e_T.05, sd_T, V)) #return a data.table with the simulated values
}Specifically:
Biological sex \((\text{S})\) is randomly sampled with replacement from a vector of factors
1(i.e., male) and2(i.e., female).Indoor air temperature \((\text{T})\) is randomly sampled from a normal distribution. The mean is set to 23°C (i.e.,
mean = 23) and the standard deviation to 2°C (i.e.,sd = 2).Clothing insulation \((\text{C})\) is randomly sampled from a truncated lognormal distribution. The mean is on the log scale (i.e.,
meanlog) and is a linear of function of biological sex \((\text{S})\) and indoor air temperature \((\text{T})\): for male (i.e.,S = 1) ismeanlog = 0.25 - 0.04*Twhile for female (i.e.,S = 2) ismeanlog = 0.25 + 0.03 - 0.04*T. The standard deviation is on the log scale and is a constant, fixed at 0.15 (i.e.,sdlog = 0.15). The parametersminandmaxare the assumed lower and upper bounds for clothing, fixed to 0.3 clo (i.e.,min = 0.3) and 2 clo (i.e.,max = 2), respectively.The measurement error \((\mathrm{e_{T}})\) is randomly sampled from a normal distribution with 0 mean (i.e.,
mean = 0) and five different values for the standard deviation (i.e.,sd = 0.1,sd = 0.2,sd = 0.3,sd = 0.4, andsd = 0.5).Thermal sensation votes \((\text{V})\) are generated from the custom function
sim_TSV.beta(), which is a function of biological sex (sex), indoor air temperature (temp), and clothing insulation (clo). The termb_intis the coefficient for the intercept (set to-8.65),b_sexis the coefficient for biological sex (set to0for males and-0.16for females),b_tempis the coefficient for indoor air temperature (set to0.37),b_clois the coefficient for clothing insulation (set to0.88), andphiis the precision parameter (set to17.9). More information on the custom functionsim_TSV.beta()can be found in Appendix A.5.
From this data generation mechanism, we simulated the target population, which consists of one million observations.
set.seed(2023) #Set random number for reproducibility
#Simulate the population
Q10_population <- data.sim_Q10.beta(N = 1e6)
#View the data frame
Q10_population S T C e_T.00 e_T.01 e_T.02 e_T.03
<fctr> <num> <num> <num> <num> <num> <num>
1: 1 19.35834 0.5173312 0 -0.062702223 -0.07056974 -0.107421688
2: 2 20.17534 0.6351932 0 0.036833817 -0.45799999 0.228275834
3: 1 17.25539 0.6350222 0 -0.036618612 0.04840667 -0.062243203
4: 1 22.32519 0.5410619 0 -0.008413221 0.32868455 -0.005429957
5: 2 23.51410 0.5319473 0 0.173145703 -0.30701565 -0.061461495
---
999996: 1 23.16175 0.5811199 0 0.096770182 0.05967175 -0.045353137
999997: 1 20.21810 0.4694916 0 -0.230831411 -0.17439661 -0.118892612
999998: 2 21.43980 0.5467828 0 -0.104000457 -0.20829030 0.255670024
999999: 1 21.73996 0.4982545 0 0.012583466 0.12697703 -0.274930898
1000000: 1 20.66438 0.6083867 0 -0.042070949 0.02382288 0.237003438
e_T.04 e_T.05 sd_T V
<num> <num> <num> <num>
1: -0.459333619 -0.07788942 0.23 0.19080604
2: 0.070525938 1.02590301 0.23 0.21231504
3: -0.129127905 0.94282810 0.23 0.01636895
4: 0.009436543 0.03922867 0.23 0.45775853
5: 0.127265614 -0.87807878 0.23 0.37499176
---
999996: -0.221818434 0.40259721 0.23 0.52735044
999997: -0.430372997 0.26680494 0.23 0.35390976
999998: 0.410357341 -0.07844362 0.23 0.43302240
999999: 0.704195878 0.38000333 0.23 0.51916711
1000000: -0.069299287 0.75154630 0.23 0.62651285From this population, we obtained a data set of ten thousand observations using simple random sampling.
set.seed(2023) #set random number for reproducibility
sample.random <-
Q10_population |>
slice_sample(n = 1e4) #take a simple random sample of size 10,000
#View the data frame
sample.random S T C e_T.00 e_T.01 e_T.02 e_T.03
<fctr> <num> <num> <num> <num> <num> <num>
1: 1 23.20618 0.4815649 0 -0.06978622 -0.129204037 0.523860078
2: 2 23.30433 0.4745407 0 -0.12912507 -0.190009463 0.057565397
3: 2 25.19966 0.6746527 0 -0.16288983 -0.172935147 0.099263168
4: 1 21.89525 0.4297182 0 -0.05043101 0.376715575 -0.264935142
5: 2 24.12164 0.4854711 0 0.09904334 0.453313611 0.159060039
---
9996: 2 21.63345 0.4495766 0 0.06705513 -0.165641646 -0.117640964
9997: 1 24.22080 0.5921024 0 0.02164574 0.138106228 0.007493248
9998: 1 23.45796 0.5325718 0 -0.04634661 -0.075558081 -0.244627137
9999: 2 22.87090 0.6053573 0 -0.03076851 0.005061156 0.203756976
10000: 1 21.96546 0.5331528 0 -0.19034684 0.122339362 -0.298962402
e_T.04 e_T.05 sd_T V
<num> <num> <num> <num>
1: -0.1553081 0.21303018 0.23 0.6571369
2: 0.6120928 0.18595653 0.23 0.5486656
3: 0.2883316 -0.46544124 0.23 0.8002563
4: -0.3662970 0.14375387 0.23 0.4289624
5: 0.4727936 -0.71967536 0.23 0.6820872
---
9996: -0.4141868 0.27983221 0.23 0.3231354
9997: -0.5735155 0.34575565 0.23 0.6442638
9998: -0.5724067 -0.03930174 0.23 0.4607735
9999: -0.5441795 0.52468814 0.23 0.5463679
10000: 0.1114434 0.18559028 0.23 0.5829363Subsequently, we added the unobserved error to the unobserved indoor air temperature \((\text{T})\) to obtain the observed indoor air temperature (\(\mathrm{T^*}\)). The unobserved error \(\mathrm{e_{T}}\) was defined as normally distributed with zero mean and five different values for the standard deviation (0.1 K, 0.2 K, 0.3 K, 0.4 K, and 0.5 K, respectively). This results in the creation of five additional data sets of ten thousand observations, each of which differs from one another only in the values of the observed indoor air temperature \((\mathrm{e_{T}})\).
Code
#No measurement error (e_T = e_T.00)
set.seed(2023) #set random number for reproducibility
sample.random00 <-
sample.random |>
mutate(T_obs = T + e_T.00,
e_T = e_T.00) |>
select(S, T, C, V, sd_T, e_T, T_obs)
#Measurement error with sd=0.1 (e_T = e_T.01)
set.seed(2023) #set random number for reproducibility
sample.random01 <-
sample.random |>
mutate(T_obs = T + e_T.01,
e_T = e_T.01) |>
select(S, T, C, V, sd_T, e_T, T_obs)
#Measurement error with sd=0.2 (e_T = e_T.02)
set.seed(2023) #set random number for reproducibility
sample.random02 <-
sample.random |>
mutate(T_obs = T + e_T.02,
e_T = e_T.02) |>
select(S, T, C, V, sd_T, e_T, T_obs)
#Measurement error with sd=0.3 (e_T = e_T.03)
set.seed(2023) #set random number for reproducibility
sample.random03 <-
sample.random |>
mutate(T_obs = T + e_T.03,
e_T = e_T.03) |>
select(S, T, C, V, sd_T, e_T, T_obs)
#Measurement error with sd=0.4 (e_T = e_T.04)
set.seed(2023) #set random number for reproducibility
sample.random04 <-
sample.random |>
mutate(T_obs = T + e_T.04,
e_T = e_T.04) |>
select(S, T, C, V, sd_T, e_T, T_obs)
#Measurement error with sd=0.5 (e_T = e_T.05)
set.seed(2023) #set random number for reproducibility
sample.random05 <-
sample.random |>
mutate(T_obs = T + e_T.05,
e_T = e_T.05) |>
select(S, T, C, V, sd_T, e_T, T_obs)We assumed that a 4-wire Pt100 resistance temperature sensing (RTD) temperature probe (tolerance class DIN B) and a transmitter that operates with a current output signal of 4 to 20 mA and a temperature range of 0 to 200°C were used for the measurement. The total measurement uncertainty (hypothetical but realistic) of the programmable transmitter with a Pt100 RTD are shown in Table 5.1.
| Source of uncertainty | Values | Probability distribution | Divisor | Standard uncertainty |
|---|---|---|---|---|
| Calibration | ± 0.025 K | Normal | 2 | ± 0.013 K |
| Thermoelectric voltage | ± 0.086 K | Rectangular | √3 | ± 0.050 K |
| Voltage supply | ± 0.050 K | Rectangular | √3 | ± 0.029 K |
| Ambient temperature | ± 0.225 K | Rectangular | √3 | ± 0.130 K |
| Linearisation and processing | ± 0.300 K | Normal | 2 | ± 0.150 K |
| Burden influence | ± 0.100 K | Rectangular | √3 | ± 0.058 K |
| Long-term stability | ± 0.100 K | Rectangular | √3 | ± 0.058 K |
The combined standard uncertainty is assumed normally distributed and is given by a quadrature sum (root of the sum of the squares of the components):
\[ \begin{aligned} u_{c}\mathrm{(T)} &= \sqrt{(0.013)^2 + (0.050)^2 + \cdots + (0.058)^2 + (0.058)^2} = 0.222 \simeq 0.23\\ \end{aligned} \]
This combined standard uncertainty is then used as standard deviation for the measurement error in air temperature (i.e., \(u_{c}\mathrm{(T)}=0.23=\sigma_{T^*}\)).
A summary of descriptive statistics for all variables in the data sets is given below. Detailed information can be found in Q.10 Summary.
summary(sample.random00) S T C V sd_T
1:4912 Min. :16.37 Min. :0.3004 Min. :0.004637 Min. :0.23
2:5088 1st Qu.:21.65 1st Qu.:0.4647 1st Qu.:0.416381 1st Qu.:0.23
Median :23.03 Median :0.5209 Median :0.559302 Median :0.23
Mean :23.00 Mean :0.5284 Mean :0.552652 Mean :0.23
3rd Qu.:24.36 3rd Qu.:0.5840 3rd Qu.:0.695347 3rd Qu.:0.23
Max. :29.98 Max. :1.0358 Max. :0.999010 Max. :0.23
e_T T_obs
Min. :0 Min. :16.37
1st Qu.:0 1st Qu.:21.65
Median :0 Median :23.03
Mean :0 Mean :23.00
3rd Qu.:0 3rd Qu.:24.36
Max. :0 Max. :29.98 summary(sample.random01) S T C V sd_T
1:4912 Min. :16.37 Min. :0.3004 Min. :0.004637 Min. :0.23
2:5088 1st Qu.:21.65 1st Qu.:0.4647 1st Qu.:0.416381 1st Qu.:0.23
Median :23.03 Median :0.5209 Median :0.559302 Median :0.23
Mean :23.00 Mean :0.5284 Mean :0.552652 Mean :0.23
3rd Qu.:24.36 3rd Qu.:0.5840 3rd Qu.:0.695347 3rd Qu.:0.23
Max. :29.98 Max. :1.0358 Max. :0.999010 Max. :0.23
e_T T_obs
Min. :-0.3803186 Min. :16.23
1st Qu.:-0.0675863 1st Qu.:21.65
Median :-0.0002152 Median :23.04
Mean :-0.0000692 Mean :23.00
3rd Qu.: 0.0668047 3rd Qu.:24.37
Max. : 0.3455248 Max. :30.08 summary(sample.random02) S T C V sd_T
1:4912 Min. :16.37 Min. :0.3004 Min. :0.004637 Min. :0.23
2:5088 1st Qu.:21.65 1st Qu.:0.4647 1st Qu.:0.416381 1st Qu.:0.23
Median :23.03 Median :0.5209 Median :0.559302 Median :0.23
Mean :23.00 Mean :0.5284 Mean :0.552652 Mean :0.23
3rd Qu.:24.36 3rd Qu.:0.5840 3rd Qu.:0.695347 3rd Qu.:0.23
Max. :29.98 Max. :1.0358 Max. :0.999010 Max. :0.23
e_T T_obs
Min. :-0.706064 Min. :16.13
1st Qu.:-0.136767 1st Qu.:21.64
Median :-0.000608 Median :23.03
Mean : 0.001003 Mean :23.00
3rd Qu.: 0.136866 3rd Qu.:24.36
Max. : 0.760917 Max. :30.00 summary(sample.random03) S T C V sd_T
1:4912 Min. :16.37 Min. :0.3004 Min. :0.004637 Min. :0.23
2:5088 1st Qu.:21.65 1st Qu.:0.4647 1st Qu.:0.416381 1st Qu.:0.23
Median :23.03 Median :0.5209 Median :0.559302 Median :0.23
Mean :23.00 Mean :0.5284 Mean :0.552652 Mean :0.23
3rd Qu.:24.36 3rd Qu.:0.5840 3rd Qu.:0.695347 3rd Qu.:0.23
Max. :29.98 Max. :1.0358 Max. :0.999010 Max. :0.23
e_T T_obs
Min. :-1.0930901 Min. :16.07
1st Qu.:-0.2027741 1st Qu.:21.64
Median :-0.0000406 Median :23.03
Mean :-0.0017450 Mean :23.00
3rd Qu.: 0.2022324 3rd Qu.:24.38
Max. : 1.1988297 Max. :30.24 summary(sample.random04) S T C V sd_T
1:4912 Min. :16.37 Min. :0.3004 Min. :0.004637 Min. :0.23
2:5088 1st Qu.:21.65 1st Qu.:0.4647 1st Qu.:0.416381 1st Qu.:0.23
Median :23.03 Median :0.5209 Median :0.559302 Median :0.23
Mean :23.00 Mean :0.5284 Mean :0.552652 Mean :0.23
3rd Qu.:24.36 3rd Qu.:0.5840 3rd Qu.:0.695347 3rd Qu.:0.23
Max. :29.98 Max. :1.0358 Max. :0.999010 Max. :0.23
e_T T_obs
Min. :-1.445697 Min. :16.16
1st Qu.:-0.278690 1st Qu.:21.59
Median :-0.009059 Median :23.03
Mean :-0.004858 Mean :23.00
3rd Qu.: 0.258943 3rd Qu.:24.38
Max. : 1.550664 Max. :30.15 summary(sample.random05) S T C V sd_T
1:4912 Min. :16.37 Min. :0.3004 Min. :0.004637 Min. :0.23
2:5088 1st Qu.:21.65 1st Qu.:0.4647 1st Qu.:0.416381 1st Qu.:0.23
Median :23.03 Median :0.5209 Median :0.559302 Median :0.23
Mean :23.00 Mean :0.5284 Mean :0.552652 Mean :0.23
3rd Qu.:24.36 3rd Qu.:0.5840 3rd Qu.:0.695347 3rd Qu.:0.23
Max. :29.98 Max. :1.0358 Max. :0.999010 Max. :0.23
e_T T_obs
Min. :-2.047073 Min. :15.68
1st Qu.:-0.327790 1st Qu.:21.63
Median : 0.004757 Median :23.03
Mean : 0.005715 Mean :23.01
3rd Qu.: 0.339385 3rd Qu.:24.40
Max. : 1.992174 Max. :31.04 5.3 Data analysis
In a scientific context, data analysis should be performed to model the phenomenon under scrutiny and not fitting curves to data. The data are a means of obtaining information about that phenomenon to answer the specific question of interest. In this example, we assumed that the objective of the data analysis is to answer the following inferential question: ‘What is the association between thermal sensation vote and sex, indoor air temperature and clothing insulation?’.
5.3.1 Scientific model
The first step to address a scientific question is to define the scientific model. A scientific model is essentially a representation of a specific phenomenon that is being studied. To achieve this, we defined a DAG. A DAG generally consists of letter-and-arrow pictures summarising the researcher’s scientific knowledge about the phenomenon under study. Since we know the data-generating process (we defined it), the scientific model equates to the data-generating process outlined in Fig. 5.1. In reality, however, it would not.2
5.3.2 Statistical model
A statistical model is a mathematical representation of the scientific model —which in turn represents the data-generating process— that aims to answer the particular question of interest. To address the question, the synthetic data sets are analysed using ordinal regression within a Bayesian framework.3
Three models are considered here:
- ‘Model 1’ is the model that uses the air temperature measured without error, \(\text{T}\). It can be considered the reference model since the resulting estimated coefficients are the ones that we would have obtained if air temperature was measured perfectly.
- ‘Model 2’ is the model that uses air temperature measured with error, \(\mathrm{T^*}\).
- ‘Model 3’ is the model that uses air temperature measured with error, \(\mathrm{T^*}\) but considering measurement error (using \(\sigma_{T^*}=0.23\)).
The statistical model for ‘Model 1’ can then be written as:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Beta}(\mu_{i}, \phi)\\ \operatorname{logit}(\mu_{i}) &= \eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{S} (\mathrm{S}_i) + \beta_{T} \mathrm{T}_{i} + \beta_{C} \mathrm{C}_{i} \\ \alpha_{0} &\sim \operatorname{Normal}(0, 5)\\ \beta_{S} &\sim \operatorname{Normal}(0, 1)\\ \beta_{T} &\sim \operatorname{Normal}(0, 1)\\ \beta_{C} &\sim \operatorname{Normal}(0, 1)\\ \phi &\sim \operatorname{Exponential}(1)\\ \end{aligned} \tag{5.1}\]
In Eq. 5.1, the conditional distribution of the response variable \(\mathrm{Y}_{i}\) for the \(i^{th}\) observation is assumed to follow a beta distribution with mean \(\mu_{i}\) and precision parameter \(\phi\). The parameter \(\eta_{i}\) is the linear predictor, where \(\alpha_{0}\) is the intercept term and \(\beta_{S}\), \(\beta_{T}\), \(\beta_{C}\) are the regression coefficients for biological sex \((\text{S})\), indoor air temperature \((\text{T})\) and clothing insulation \((\text{C})\), respectively. The intercept \(\alpha_{0}\) has a \(\operatorname{Normal}(0, 5)\) prior, the regression coefficients \(\beta_{S}\), \(\beta_{T}\), \(\beta_{C}\) have a \(\operatorname{Normal}(0, 1)\) prior, and the precision parameter \(\phi\) has an \(\operatorname{Exponential}(1)\) prior.
The statistical model for ‘Model 2’ can then be written as:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Beta}(\mu_{i}, \phi)\\ \operatorname{logit}(\mu_{i}) &= \eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{S} (\mathrm{S}_i) + \beta_{T^*} \mathrm{T}{^*_i} + \beta_{C} \mathrm{C}_{i} \\ \alpha_{0} &\sim \operatorname{Normal}(0, 5)\\ \beta_{S} &\sim \operatorname{Normal}(0, 1)\\ \beta_{T^*} &\sim \operatorname{Normal}(0, 1)\\ \beta_{C} &\sim \operatorname{Normal}(0, 1)\\ \phi &\sim \operatorname{Exponential}(1)\\ \end{aligned} \tag{5.2}\] In Eq. 5.2, the only difference from Eq. 5.1 is the presence of \(\mathrm{T}{^*}\) instead of \(\text{T}\) in the linear predictor \(\eta_{i}\). The prior for the regression coefficient \(\beta_{T^*}\) is equal to the prior of \(\beta_{T}\) (i.e., \(\operatorname{Normal}(0, 1)\)).
The statistical model for ‘Model 3’ can then be written as:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Beta}(\mu_{i}, \phi)\\ \operatorname{logit}(\mu_{i}) &= \eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{S} (\mathrm{S}_i) + \beta_{T} \mathrm{T}_{i} + \beta_{C} \mathrm{C}_{i}\\ \mathrm{T}{^*_i} &\sim \operatorname{Normal}(\mathrm{T}_{i}, \sigma_{T^*})\\ \mathrm{T}_{i} &\sim \operatorname{Normal}(\mu_{T}, \sigma_{T})\\ \alpha_{0} &\sim \operatorname{Normal}(0, 5)\\ \beta_{S} &\sim \operatorname{Normal}(0, 1)\\ \beta_{T} &\sim \operatorname{Normal}(0, 1)\\ \beta_{C} &\sim \operatorname{Normal}(0, 1)\\ \mu_{T} &\sim \operatorname{Normal}(22, 7)\\ \sigma_{T} &\sim \operatorname{Exponential}(1)\\ \phi &\sim \operatorname{Exponential}(1)\\ \end{aligned} \tag{5.3}\]
In Eq. 5.3, the terms in the linear predictor \(\eta_{i}\) are the same as in Eq. 5.1, with a difference: \(\text{T}\) is now declared as a parameter rather than as data. The true value \(\text{T}\) is treated as missing data and estimated along with other quantities in the model. The data are now \(\mathrm{T}{^*_i}\), which are modelled as normally distributed with mean \(\mathrm{T}{_i}\) and standard deviation \(\sigma_{T^*}\) (i.e., \(\operatorname{Normal}(\mathrm{T}_{i}, \sigma_{T^*})\)). Furthermore, the true values \(\mathrm{T}{_i}\) are given a hierarchical prior (i.e., \(\operatorname{Normal}(\mu_{T}, \sigma_{T})\)), assuming that each \(\mathrm{T}{_i}\) is normally distributed with an underlying mean \(\mu_{T}\), and a standard deviation \(\sigma_{T}\). The latent mean \(\mu_{T}\) has a \(\operatorname{Normal}(22, 7)\) prior, and the standard deviation \(\sigma_{T}\) has a \(\operatorname{Exponential}(1)\) prior. The parameters \(\mu_{T}\) and \(\sigma_{T}\) are often referred to as hyperparameters (i.e., they are parameters for parameters) and thier priors (i.e., \(\operatorname{Normal}(22, 7)\) and \(\operatorname{Exponential}(1)\)) are often called hyperpriors.
A summary description of the simulation example is provided in Table 5.2.
| Pitfall | Ignore the implications of measurement error on the results and interpretation of a data analysis. |
| Type of analysis | Inferential. |
| Research question | What is the association between TSV and sex, indoor air temperature and clothing insulation? |
| Framework | Bayesian. |
| Assumptions | Random sample (simple random sampling): everyone in the population has an equal chance of being selected into the sample. |
| Limited random variability: large sample size. | |
| Independence of observations: each observation represents independent bits of information. | |
| No confounding: the DAG includes all shared causes among the variables. | |
| No model error: perfect functional form specification. | |
| Variables | Unobserved indoor air temperature (T): continuous variable [unit: °C] |
| Observed indoor air temperature (T*): continuous variable [unit: °C] | |
| Thermal resistance of clothing (C): continuous variable [unit: clo] | |
| Sex (S): categorical variable [‘1’ male; ‘2’ female] | |
| TSV (V): continuous but bounded variable with interval (0, 1) [‘0’ cold; ‘1’ hot] |
5.3.3 Results
The results of the fitted statistical models (defined above) are presented here. There a total of 13 fitted models:
- Three models for the data set with no measurement error (i.e.,
Model.1_e00,Model.2_e00andModel.3_e00). - Two models for the data set with measurement error equal to 0.1 standard deviation (i.e.,
Model.2_e01andModel.3_e01). - Two models for the data set with measurement error equal to 0.2 standard deviation (i.e.,
Model.2_e02andModel.3_e02). - Two models for the data set with measurement error equal to 0.3 standard deviation (i.e.,
Model.2_e03andModel.3_e03). - Two models for the data set with measurement error equal to 0.4 standard deviation (i.e.,
Model.2_e04andModel.3_e04). - Two models for the data set with measurement error equal to 0.5 standard deviation (i.e.,
Model.2_e05andModel.3_e05).
brms syntax for the formula argument
In brms, the intercept can be specified by the syntax y ~ x or y ~ 1 + x in the formula argument. When doing so the corresponding prior is applied under the presupposition all the regressors (x) are mean centred. However, if the regressors are not mean centred (like in our case), the syntax ... ~ 0 + Intercept + ... should be used. With this syntax, the prior can be defined on the real intercept, directly. More information can be found in the brms reference manual.
Code
#No measurement error (e_T = e_T.00)
Model.1_e00 <-
brm(data = sample.random00,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q10_mod.1_00') #save model fit
Model.2_e00 <-
brm(data = sample.random00,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T_obs + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q10_mod.2_00') #save model fit
Model.3_e00 <-
brm(data = sample.random00,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + me(T_obs, sd_T) + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = meT_obssd_T),
prior(normal(22, 7), class = meanme, coef = meT_obs),
prior(exponential(1), class = sdme, coef = meT_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
#Save the draws of latent variables obtained by using the 'me' term (default is FALSE)
save_pars = save_pars(latent = TRUE),
file = 'fit/Q10_mod.3_00') #save model fit
#Measurement error with sd=0.1 (e_T = e_T.01)
Model.2_e01 <-
brm(data = sample.random01,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T_obs + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q10_mod.2_01') #save model fit
Model.3_e01 <-
brm(data = sample.random00,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + me(T_obs, sd_T) + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = meT_obssd_T),
prior(normal(22, 7), class = meanme, coef = meT_obs),
prior(exponential(1), class = sdme, coef = meT_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
#Save the draws of latent variables obtained by using the 'me' term (default is FALSE)
save_pars = save_pars(latent = TRUE),
file = 'fit/Q10_mod.3_01') #save model fit
#Measurement error with sd=0.2 (e_T = e_T.02)
Model.2_e02 <-
brm(data = sample.random02,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T_obs + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q10_mod.2_02') #save model fit
Model.3_e02 <-
brm(data = sample.random02,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + me(T_obs, sd_T) + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = meT_obssd_T),
prior(normal(22, 7), class = meanme, coef = meT_obs),
prior(exponential(1), class = sdme, coef = meT_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
#Save the draws of latent variables obtained by using the 'me' term (default is FALSE)
save_pars = save_pars(latent = TRUE),
file = 'fit/Q10_mod.3_02') #save model fit
#Measurement error with sd=0.3 (e_T = e_T.03)
Model.2_e03 <-
brm(data = sample.random03,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T_obs + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q10_mod.2_03') #save model fit
Model.3_e03 <-
brm(data = sample.random03,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + me(T_obs, sd_T) + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = meT_obssd_T),
prior(normal(22, 7), class = meanme, coef = meT_obs),
prior(exponential(1), class = sdme, coef = meT_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
#Save the draws of latent variables obtained by using the 'me' term (default is FALSE)
save_pars = save_pars(latent = TRUE),
file = 'fit/Q10_mod.3_03') #save model fit
#Measurement error with sd=0.4 (e_T = e_T.04)
Model.2_e04 <-
brm(data = sample.random04,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T_obs + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q10_mod.2_04') #save model fit
Model.3_e04 <-
brm(data = sample.random04,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + me(T_obs, sd_T) + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = meT_obssd_T),
prior(normal(22, 7), class = meanme, coef = meT_obs),
prior(exponential(1), class = sdme, coef = meT_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
#Save the draws of latent variables obtained by using the 'me' term (default is FALSE)
save_pars = save_pars(latent = TRUE),
file = 'fit/Q10_mod.3_04') #save model fit
#Measurement error with sd=0.5 (e_T = e_T.05)
Model.2_e05 <-
brm(data = sample.random05,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T_obs + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q10_mod.2_05') #save model fit
Model.3_e05 <-
brm(data = sample.random05,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + me(T_obs, sd_T) + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = meT_obssd_T),
prior(normal(22, 7), class = meanme, coef = meT_obs),
prior(exponential(1), class = sdme, coef = meT_obs),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
#Save the draws of latent variables obtained by using the 'me' term (default is FALSE)
save_pars = save_pars(latent = TRUE),
file = 'fit/Q10_mod.3_05') #save model fitSubsequently, the posterior distributions for the 13 models are extracted and summarised.
Code
#Extract posterior and summarise
#Models with no measurement error (e_T = e_T.00)
post.mod.1_00 <-
Model.1_e00 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, b_T) |> #select 'b_S2', 'b_C', and 'b_T'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = b_T) |> #rename 'b_T' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.1', #create a column
Error = '0.0') #create a column
post.mod.2_00 <-
Model.2_e00 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, b_T_obs) |> #select 'b_S2', 'b_C', and 'b_T_obs'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = b_T_obs) |> #rename 'b_T_obs' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.2', #create a column
Error = '0.0') #create a column
post.mod.3_00 <-
Model.3_e00 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, bsp_meT_obssd_T) |> #select 'b_S2', 'b_C', and 'bsp_meT_obssd_T'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = bsp_meT_obssd_T) |> #rename 'bsp_meT_obssd_T' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.3', #create a column
Error = '0.0') #create a column
#Models with measurement error with sd=0.1 (e_T = e_T.01)
post.mod.2_01 <-
Model.2_e01 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, b_T_obs) |> #select 'b_S2', 'b_C', and 'b_T_obs'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = b_T_obs) |> #rename 'b_T_obs' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.2', #create a column
Error = '0.1') #create a column
post.mod.3_01 <-
Model.3_e01 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, bsp_meT_obssd_T) |> #select 'b_S2', 'b_C', and 'bsp_meT_obssd_T'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = bsp_meT_obssd_T) |> #rename 'bsp_meT_obssd_T' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.3', #create a column
Error = '0.1') #create a column
#Models with measurement error with sd=0.2 (e_T = e_T.02)
post.mod.2_02 <-
Model.2_e02 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, b_T_obs) |> #select 'b_S2', 'b_C', and 'b_T_obs'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = b_T_obs) |> #rename 'b_T_obs' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.2', #create a column
Error = '0.2') #create a column
post.mod.3_02 <-
Model.3_e02 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, bsp_meT_obssd_T) |> #select 'b_S2', 'b_C', and 'bsp_meT_obssd_T'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = bsp_meT_obssd_T) |> #rename 'bsp_meT_obssd_T' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.3', #create a column
Error = '0.2') #create a column
#Models with measurement error with sd=0.3 (e_T = e_T.03)
post.mod.2_03 <-
Model.2_e03 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, b_T_obs) |> #select 'b_S2', 'b_C', and 'b_T_obs'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = b_T_obs) |> #rename 'b_T_obs' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.2', #create a column
Error = '0.3') #create a column
post.mod.3_03 <-
Model.3_e03 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, bsp_meT_obssd_T) |> #select 'b_S2', 'b_C', and 'bsp_meT_obssd_T'
rename(Sex = b_S2, #rename 'b_S2' as Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = bsp_meT_obssd_T) |> #rename 'bsp_meT_obssd_T' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.3', #create a column
Error = '0.3') #create a column
#Models with measurement error with sd=0.4 (e_T = e_T.04)
post.mod.2_04 <-
Model.2_e04 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, b_T_obs) |> #select 'b_S2', 'b_C', and 'b_T_obs'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = b_T_obs) |> #rename 'b_T_obs' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.2', #create a column
Error = '0.4') #create a column
post.mod.3_04 <-
Model.3_e04 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, bsp_meT_obssd_T) |> #select 'b_S2', 'b_C', and 'bsp_meT_obssd_T'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = bsp_meT_obssd_T) |> #rename 'bsp_meT_obssd_T' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.3', #create a column
Error = '0.4') #create a column
#Models with measurement error with sd=0.5 (e_T = e_T.05)
post.mod.2_05 <-
Model.2_e05 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, b_T_obs) |> #select 'b_S2', 'b_C', and 'b_T_obs'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = b_T_obs) |> #rename 'b_T_obs' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.2', #create a column
Error = '0.5') #create a column
post.mod.3_05 <-
Model.3_e05 |>
as_draws_df() |> #transform a brmsfit object to a format supported by the posterior package
data.table() |> #convert to data.table format
select(b_S2, b_C, bsp_meT_obssd_T) |> #select 'b_S2', 'b_C', and 'bsp_meT_obssd_T'
rename(Sex = b_S2, #rename 'b_S2' as 'Sex'
Clothing = b_C, #rename 'b_C' as 'Clothing'
Air.Temperature = bsp_meT_obssd_T) |> #rename 'bsp_meT_obssd_T' as 'Air.Temperature'
pivot_longer(c(Sex, Clothing, Air.Temperature), #convert to the long format
names_to = 'Coef', #add columns name here
values_to = 'Value') |> #add values here
group_by(Coef) |> #group by 'Coef'
mean_hdi(Value) |> #point (mean) and interval (Highest Density Intervals, HDI) summaries of the coefficients' distribution
mutate(Model = 'mod.3', #create a column
Error = '0.5') #create a column
#Combine in one data frame
post <- data.table(rbind(post.mod.1_00, post.mod.2_00, post.mod.3_00,
post.mod.2_01, post.mod.3_01,
post.mod.2_02, post.mod.3_02,
post.mod.2_03, post.mod.3_03,
post.mod.2_04, post.mod.3_04,
post.mod.2_05, post.mod.3_05))
#Transform 'Model' to an ordered factor
post[, Model := factor(Model, levels = c('mod.3','mod.2','mod.1'), ordered = TRUE)]
#Transform 'Coef' to an ordered factor
post[, Coef := factor(Coef, levels = c('Sex', 'Air.Temperature', 'Clothing'), ordered = TRUE)]The results are then plotted in Fig. 5.2.
Code
coef.sim <-
data.table(Coef = unique(post$Coef), Value = c(0.37, 0.88, -0.16))
Coef_names <- c('Intercept' = 'Intercepet',
'Sex' = 'Sex',
'Air.Temperature' = 'Air Temperature',
'Clothing' = 'Clothing')
Error_names <- c('0.0' = 'No error',
'0.1' = 'sd = 0.1',
'0.2' = 'sd = 0.2',
'0.3' = 'sd = 0.3',
'0.4' = 'sd = 0.4',
'0.5' = 'sd = 0.5')
ggplot(data = post, aes(x = Value, y = Model, xmin = .lower, xmax = .upper, colour = Model)) +
geom_vline(data = coef.sim, aes(xintercept = Value),
linetype = 'solid',
linewidth = 0.4,
colour = 'grey60') +
geom_linerange(linewidth = 0.75) +
geom_point(size = 1.5) +
geom_text(aes(x = Value, y = Model, label = paste(sprintf('%.3f', Value))),
size = 5.5/.pt, colour = 'black', vjust = -0.9, hjust = 0.5) +
geom_text(aes(x = .lower, y = Model, label = paste(sprintf('%.2f', .lower))),
size = 5.5/.pt, colour = 'black', vjust = -0.7, hjust = 0.6) +
geom_text(aes(x = .upper, y = Model, label = paste(sprintf('%.2f', .upper))),
size = 5.5/.pt, colour = 'black', vjust = -0.7, hjust = 0.4) +
scale_colour_manual('Models', values = c('black', '#F65314', '#7CBB00'),
breaks=c('mod.1', 'mod.2', 'mod.3'),
labels = c('Model 1', 'Model 2', 'Model 3')) +
scale_x_continuous('Estimate of the coefficients',
breaks = coef.sim$Value) +
facet_grid(Error ~ Coef, scales = 'free',
labeller = labeller(Error = Error_names,
Coef = Coef_names)) +
theme(legend.position = 'bottom',
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
axis.title.y = element_blank(),
axis.text.y = element_blank(),
axis.ticks.y = element_blank(),
axis.title.x = element_text(margin = unit(c(2, 0, 0, 0), "mm"))
)
Fig. 5.2 shows the estimates of the regression coefficients for three models and their 95% credible intervals. Here, the estimated posterior distributions of the regression coefficients for ‘Model 1’, ‘Model 2’ and ‘Model 3’ are shown by the black, orange and green lines, respectively. With regard to the increment in measurement error (i.e., from ‘No error’ to sd=0.5), the following comparison can be made:
- The bias of the sex coefficient is positive and very similar in magnitude for both ‘Model 2’ and ‘Model 3’;
- The bias of the temperature coefficient is negative (i.e., regression dilution) for ‘Model 2’ and ‘Model 3’ but overall higher in magnitude for ‘Model 2’. However, in ‘Model 3’ considering measurement error when it does not exist (i.e., ‘no error’) or is lower in magnitude than the assumed one (i.e., \(\sigma_{T^*}=0.23>(\text{sd}=0.1~\text{and}~\text{sd}=0.2)\)) undermines the inference by overestimating the coefficient;
- The bias of the clothing coefficient is negative and very similar in magnitude for both ‘Model 2’ and ‘Model 3’.
Considering measurement error in statistical modelling is essential, but it is important to proceed cautiously as no ‘one-size-fits-all’ solutions are applicable. The presence of measurement error can lead to unpredictable effects; it can over- or underestimate the coefficients of interest depending on which variables are affected, how the measurement error is structured and its magnitude. Although directly modelling measurement error can be complicated, evaluating its implications for the results and conclusions is always possible by performing a sensitivity analysis.
Session info
Version information about R, the OS and attached or loaded packages.
R version 4.2.3 (2023-03-15 ucrt)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 22631)
Matrix products: default
locale:
[1] LC_COLLATE=English_United Kingdom.utf8
[2] LC_CTYPE=English_United Kingdom.utf8
[3] LC_MONETARY=English_United Kingdom.utf8
[4] LC_NUMERIC=C
[5] LC_TIME=English_United Kingdom.utf8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] EnvStats_2.8.1 brms_2.21.0 Rcpp_1.0.12 tidybayes_3.0.6
[5] lubridate_1.9.3 forcats_1.0.0 stringr_1.5.1 dplyr_1.1.4
[9] purrr_1.0.2 readr_2.1.5 tidyr_1.3.1 tibble_3.2.1
[13] tidyverse_2.0.0 gt_0.11.0 data.table_1.15.4 ggplot2_3.5.1
[17] dagitty_0.3-4 ggdag_0.2.12
loaded via a namespace (and not attached):
[1] nlme_3.1-162 matrixStats_1.3.0 rstan_2.32.6
[4] tensorA_0.36.2.1 tools_4.2.3 backports_1.4.1
[7] utf8_1.2.4 R6_2.5.1 colorspace_2.1-0
[10] ggdist_3.3.2 withr_3.0.0 tidyselect_1.2.1
[13] gridExtra_2.3 Brobdingnag_1.2-9 emmeans_1.10.3
[16] curl_5.2.1 compiler_4.2.3 cli_3.6.2
[19] arrayhelpers_1.1-0 xml2_1.3.6 sass_0.4.9
[22] labeling_0.4.3 posterior_1.6.0 scales_1.3.0
[25] checkmate_2.3.1 mvtnorm_1.2-4 QuickJSR_1.3.1
[28] digest_0.6.36 StanHeaders_2.32.10 rmarkdown_2.27
[31] pkgconfig_2.0.3 htmltools_0.5.8.1 fastmap_1.2.0
[34] htmlwidgets_1.6.4 rlang_1.1.4 rstudioapi_0.16.0
[37] farver_2.1.2 generics_0.1.3 svUnit_1.0.6
[40] jsonlite_1.8.8 distributional_0.4.0 inline_0.3.19
[43] magrittr_2.0.3 loo_2.8.0 bayesplot_1.11.1
[46] Matrix_1.6-5 munsell_0.5.1 fansi_1.0.6
[49] viridis_0.6.5 abind_1.4-5 lifecycle_1.0.4
[52] stringi_1.8.4 yaml_2.3.9 ggraph_2.2.1
[55] MASS_7.3-58.2 pkgbuild_1.4.4 grid_4.2.3
[58] ggrepel_0.9.5 parallel_4.2.3 lattice_0.20-45
[61] graphlayouts_1.1.1 hms_1.1.3 knitr_1.48
[64] pillar_1.9.0 igraph_2.0.3 boot_1.3-28.1
[67] estimability_1.5.1 codetools_0.2-19 stats4_4.2.3
[70] rstantools_2.4.0 glue_1.7.0 evaluate_0.24.0
[73] V8_4.4.2 RcppParallel_5.1.8 tweenr_2.0.3
[76] vctrs_0.6.5 tzdb_0.4.0 polyclip_1.10-6
[79] gtable_0.3.5 cachem_1.1.0 ggforce_0.4.2
[82] xfun_0.46 xtable_1.8-4 tidygraph_1.3.1
[85] coda_0.19-4.1 viridisLite_0.4.2 memoise_2.0.1
[88] timechange_0.3.0 bridgesampling_1.1-2We have selected only a few variables of interest to limit the complexity of the DAG and shift the focus to statistical data analysis.↩︎
The scientific model is generally an approximation (a best guess) of the unknown data-generating process. As such, it would have some inaccuracies/errors, which will inevitably affect the data analysis.↩︎
In all simulation examples, we relied on the regression framework (ordinal or beta regression) within the frequentist or Bayesian approach. However, other approaches are available and could be used depending on the question and the data. It is the researcher’s responsibility to justify the choice of the statistical framework (and any decisions made during the modelling process) to answer the specific question of interest.↩︎