4 Simulation example Q.9
In this simulation example, the highlighted data analysis pitfall is clustered/nested observations. Specifically, it shows the implication of having clustered/nested observations on the result of the data analysis.
4.1 Data-generating process
The data-generating process is the specific process that generates the observed data. In reality, this process is unknown and understanding it is, in a general sense, the ultimate aim of science. Here, we assume this process is known and describe it via a directed acyclic graph (DAG) in Fig. 4.1. In this DAG,1 indoor air temperature \((\text{T})\) and biological sex \((\text{S})\) influence thermal sensation vote \((\text{V})\) both directly and indirectly, passing through clothing insulation \((\text{C})\). In addition, thermal sensation vote \((\text{V})\) is influenced directly by some unmeasured characteristics of the specific building \((\text{B})\).
Code
dag_coords <-
data.table(name = c('T', 'C', 'S', 'V', 'B'),
x = c(1, 2, 3, 2, 1),
y = c(1.2, 1.2, 1.2, 1, 1))
DAG.09 <- dagify(C ~ S,
C ~ T,
V ~ S + C + T + B,
coords = dag_coords)
ggplot(data = DAG.09, aes(x = x, y = y, xend = xend, yend = yend)) +
geom_dag_text(colour='black', size = 8, parse = TRUE,
family = c('mono'),
label = c(expression(bold(B)), expression(bold(C)), expression(bold(S)), expression(bold(T)), expression(bold(V)))) +
geom_dag_edges(arrow_directed = grid::arrow(length = grid::unit(8, 'pt'), type = 'open'),
family = c('mono'),
fontface = c('bold')) +
coord_cartesian(xlim = c(1, 3), ylim = c(1, 1.2)) +
theme_dag()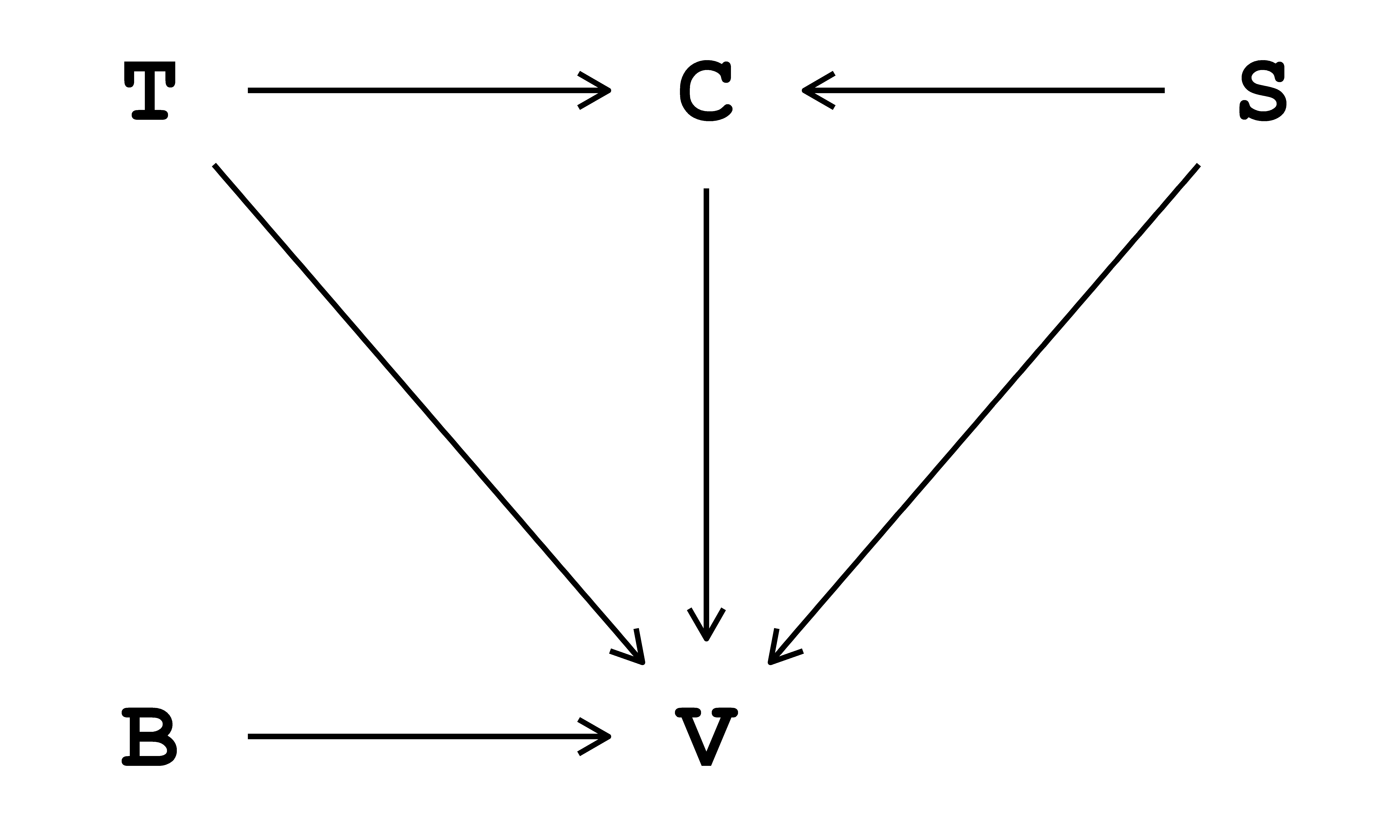
The DAG in Fig. 4.1 can be written as:
- \(C \sim f_{C}(S, T)\), read as ‘clothing insulation \((\text{C})\) is some function of biological sex \((\text{S})\) and indoor air temperature \((\text{T})\)’.
- \(V \sim f_{V}(S, T, C, B)\), read as ‘thermal sensation vote \((\text{V})\) is some function of biological sex \((\text{S})\), indoor air temperature \((\text{T})\), clothing insulation \((\text{C})\) and some unmeasured characteristics of the building \((\text{B})\)’.
4.2 Synthetic data set
In a DAG, an omitted arrow between two variables signifies the absence of direct influence of one variable on another. In contrast, a present arrow does not provide any information about the magnitude of the effect or its functional form. As such, all relationships are qualitative. However, to generate synthetic data, we need to quantitatively define the relationship between the variables. These relationships are defined within the custom function data.sim_Q09.beta(). This function takes as input the sample size N and the number of buildings n_group, and generates synthetic data according to the DAG in Fig. 4.1.
data.sim_Q09.beta <- function(N, n_group) {
#Simulate the number of people in each of the ngroup buildings so that the total number of people is N.
B_size <- rmultinom(n = 1, size=N, prob = sample(c(0.1), size=n_group, replace = TRUE))[,1]
#Simulate the building random effects
B_B <- rnorm(n_group, mean=0 , sd=0.75)
#Simulate indoor air temperature for each building
T_B <- rnorm(n_group, mean=23 , sd=2)
#Create empty list
listID <- list()
listB <- list()
listT <- list()
#Replicate group-level variables (e.g., indoor air temperature) for each observations
for (i in 1:n_group){
listB[[i]] <- rep(B_B[i], times=B_size[i]) #replicate B_B[i] for B_size[i] times
listID[[i]] <- rep((1:n_group)[i], times=B_size[i]) #replicate (1:ngroup)[i] for B_size[i] times
listT[[i]] <- rep(T_B[i], times=B_size[i]) #replicate T_B[i] for B_size[i] times
}
#Save the lists as vectors
B_id <- unlist(listID) #a vector of building identifiers of size N
B <- unlist(listB) #a vector of building random effects of size N
T <- unlist(listT) #a vector of indoor air temperature of size N
#Simulate biological sex
S <- factor(sample(1:2, size=N, replace=TRUE)) #S = 1 'male'; S = 2 'female'
#Simulate clothing insulation
C <- rlnormTrunc(N, meanlog = ifelse(S=='1', 0.25 + 0.00 -0.04*T, 0.25 + 0.03 -0.04*T), sdlog = 0.15, min = 0.3 , max = 2)
#Simulate thermal sensation vote
V <- sim_TSV.beta(n=N, sex=S, temp=T, clo=C, b_int=-8.65, b_sex=c(0,-0.16), b_temp=0.37, b_clo=0.88, build=B, phi=17.9)
return(data.table(S, T, C, V, B, B_id)) #return a data.table with the simulated values
}Specifically:
The buildings random effect \((\text{B})\) are randomly sampled from a normal distribution with 0 mean (i.e.,
mean = 0) and 0.75 standard deviation (i.e.,sd = 0.75).Indoor air temperature \((\text{T})\) is randomly sampled from a normal distribution. The mean is set to 23°C (i.e.,
mean = 23) and the standard deviation to 2°C (i.e.,sd = 2). Indoor air temperature is simulated as a between-group variable, assuming that each building has one indoor air temperature.Biological sex \((\text{S})\) is randomly sampled with replacement from a vector of factors
1(i.e., male) and2(i.e., female). Biological sex is simulated as a within-group variable, assuming each observation has a value independent from the specific building.Clothing insulation \((\text{C})\) is randomly sampled from a truncated lognormal distribution. The mean is on the log scale (i.e.,
meanlog) and is a linear of function of biological sex \((\text{S})\) and indoor air temperature \((\text{T})\): for male (i.e.,S = 1) ismeanlog = 0.25 + 0.00 -0.04*Twhile for female (i.e.,S = 2) ismeanlog = 0.25 + 0.03 -0.04*T. The standard deviation is on the log scale and is a constant, fixed at 0.15 (i.e.,sdlog = 0.15). The parametersminandmaxare the assumed lower and upper bounds for clothing, fixed to 0.3 clo (i.e.,min = 0.3) and 2 clo (i.e.,max = 2), respectively. Clothing insulation is simulated as a within-group variable, assuming each observation has a value independent from the specific building.Thermal sensation votes \((\text{V})\) are generated from the custom function
sim_TSV.beta(), which is a function of biological sex (sex), indoor air temperature (temp), and clothing insulation (clo). The termb_intis the coefficient for the intercept (set to-8.65),b_sexis the coefficient for biological sex (set to0for males and-0.16for females),b_tempis the coefficient for indoor air temperature (set to0.37),b_clois the coefficient for clothing insulation (set to0.88), andphiis the precision parameter (set to17.9). More information on the custom functionsim_TSV.beta()can be found in Appendix A.5.
From this data generation mechanism, we simulated the target population, which consists of one million observations from twenty thousand buildings.
set.seed(2023) #set random number for reproducibility
#Simulate the population
Q09_population <- data.sim_Q09.beta(N = 1e6, n_group = 20000)
#View the data frame
Q09_population S T C V B B_id
<fctr> <num> <num> <num> <num> <int>
1: 2 22.81022 0.4900057 0.8272881 1.2676635 1
2: 2 22.81022 0.3467289 0.9028027 1.2676635 1
3: 1 22.81022 0.4034637 0.5864774 1.2676635 1
4: 2 22.81022 0.5417691 0.6345086 1.2676635 1
5: 1 22.81022 0.5495862 0.7164846 1.2676635 1
---
999996: 2 27.34685 0.4707264 0.8875221 0.6904279 20000
999997: 1 27.34685 0.4239501 0.9774967 0.6904279 20000
999998: 1 27.34685 0.4790684 0.9720889 0.6904279 20000
999999: 1 27.34685 0.3614063 0.9445903 0.6904279 20000
1000000: 2 27.34685 0.3819760 0.7213163 0.6904279 20000From this population, we obtain a data set of ten thousand observations using multistage sampling. We first randomly sampled 500 buildings; among the sampled buildings, we randomly sampled ten thousand observations.
set.seed(2023) #Set random number for reproducibility
#Sample a vector of 500 unique buildings ID
sample.building <-
Q09_population$B_id |> #select the building ID
unique() |> #remove duplicate values
sample(size = 5e2) #take a simple random sample of size 500
set.seed(2023) #set random number for reproducibility
#Sample one data set of 10,000 observations
sample.random <-
Q09_population |>
filter(B_id %in% sample.building) |> #filter by the 500 building ID
slice_sample(n = 1e4) #take a simple random sample of size 10,000 from the 500 buildings
#View the data frame
sample.random S T C V B B_id
<fctr> <num> <num> <num> <num> <int>
1: 1 26.22338 0.4057381 0.4840411 -1.2220967 16952
2: 1 25.19498 0.4276205 0.7277264 -0.2434178 8566
3: 1 19.41889 0.5108458 0.2018259 0.4214767 1467
4: 2 22.85782 0.3410838 0.5978592 0.6718118 6646
5: 2 23.16726 0.4505068 0.5852178 -0.1235028 6518
---
9996: 2 19.15630 0.7439215 0.3920396 0.4903466 10025
9997: 2 19.85480 0.5591411 0.1787886 -0.5343069 7528
9998: 2 24.89526 0.5089781 0.7077378 0.5052873 11047
9999: 1 22.17644 0.4353788 0.6363981 0.2581817 15913
10000: 1 23.58039 0.4962462 0.9294576 0.4319900 9757A summary of descriptive statistics for all variables in the data sets is given below. Detailed information can be found in Q.9 Summary.
summary(sample.random) S T C V
1:4971 Min. :18.13 Min. :0.3015 Min. :0.005116
2:5029 1st Qu.:21.85 1st Qu.:0.4613 1st Qu.:0.377474
Median :23.23 Median :0.5167 Median :0.569528
Mean :23.18 Mean :0.5246 Mean :0.559469
3rd Qu.:24.46 3rd Qu.:0.5803 3rd Qu.:0.749002
Max. :28.83 Max. :1.0351 Max. :0.999927
B B_id
Min. :-2.21596 Min. : 2
1st Qu.:-0.54383 1st Qu.: 5063
Median : 0.01985 Median : 9920
Mean :-0.01413 Mean : 9951
3rd Qu.: 0.51081 3rd Qu.:14998
Max. : 2.10181 Max. :19887 4.3 Data analysis
In a scientific context, data analysis should be performed to model the phenomenon under scrutiny and not fitting curves to data. The data are a means of obtaining information about that phenomenon to answer the specific question of interest. In this example, we assumed that the objective of the data analysis is to answer the following predictive question: ‘What is the probability that the thermal sensation vote for females at 25°C with a clothing insulation value of 0.6 clo in a typical (i.e., average) building is lower than 0.5 (‘neutral’)?’.
4.3.1 Scientific model
The first step to address a scientific question is to define the scientific model. A scientific model is essentially a representation of a specific phenomenon that is being studied. To achieve this, we defined a DAG. A DAG generally consists of letter-and-arrow pictures summarising the researcher’s scientific knowledge about the phenomenon under study. Since we know the data-generating process (we defined it), the scientific model equates to the data-generating process outlined in Fig. 4.1. In reality, however, it would not.2
4.3.2 Statistical model
A statistical model is a mathematical representation of the scientific model —which in turn represents the data-generating process— that aims to answer the particular question of interest. To address the question, the synthetic data sets are analysed using beta regression within a Bayesian framework.3
To estimate the effect of interest, two beta regression models are fitted: one that ignores the cluster present in the data (Model 1) and one that considers it (Model 2).
The statistical model for ‘Model 1’ can then be written as:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Beta}(\mu_{i}, \phi)\\ \operatorname{logit}(\mu_{i}) &= \eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{S} (\mathrm{S}_i) + \beta_{T} \mathrm{T}_{i} + \beta_{C} \mathrm{C}_{i} \\ \alpha_{0} &\sim \operatorname{Normal}(0, 5)\\ \beta_{S} &\sim \operatorname{Normal}(0, 1)\\ \beta_{T} &\sim \operatorname{Normal}(0, 1)\\ \beta_{C} &\sim \operatorname{Normal}(0, 1)\\ \phi &\sim \operatorname{Exponential}(1)\\ \end{aligned} \tag{4.1}\]
In Eq. 5.1, the conditional distribution of the response variable \(\mathrm{Y}_{i}\) for the \(i^{th}\) observation is assumed to follow a beta distribution with mean \(\mu_{i}\) and precision parameter \(\phi\). The parameter \(\eta_{i}\) is the linear predictor, where \(\alpha_{0}\) is the intercept term and \(\beta_{S}\), \(\beta_{T}\), \(\beta_{C}\) are the regression coefficients for biological sex \((\text{S})\), indoor air temperature \((\text{T})\) and clothing insulation \((\text{C})\), respectively. The intercept \(\alpha_{0}\) has a \(\operatorname{Normal}(0, 5)\) prior, the regression coefficients \(\beta_{S}\), \(\beta_{T}\), \(\beta_{C}\) have a \(\operatorname{Normal}(0, 1)\) prior, and the precision parameter \(\phi\) has an \(\operatorname{Exponential}(1)\) prior.
The statistical model for ‘Model 2’ can then be written as:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Beta}(\mu_{i}, \phi)\\ \operatorname{logit}(\mu_{i}) &= \eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{S} (\mathrm{S}_i) + \beta_{T} \mathrm{T}_{i} + \beta_{C} \mathrm{C}_{i} + \alpha_{\mathrm{BUILD}[i]} \\ \alpha_{0} &\sim \operatorname{Normal}(0, 5)\\ \beta_{S} &\sim \operatorname{Normal}(0, 1)\\ \beta_{T} &\sim \operatorname{Normal}(0, 1)\\ \beta_{C} &\sim \operatorname{Normal}(0, 1)\\ \alpha_{j} &\sim \operatorname{Normal}(0, 1) ~\text{for building}~j=1 .. 500\\ \phi &\sim \operatorname{Exponential}(1)\\ \end{aligned} \tag{4.2}\]
In Eq. 5.2, the only difference from Eq. 5.1 is the presence of \(\alpha_{\mathrm{BUILD}[i]}\) in the linear predictor \(\eta_{i}\). The parameter \(\alpha_{j}\) is the random effect (specifically a random intercept) that accounts for the specific effect of the \(j^{th}\) building \(\mathrm{B}_{j}\), assuming a \(\operatorname{Normal}(0, 1)\) prior.
A summary description of the simulation example is provided in Table 4.1.
| Pitfall | Ignore the implications of clustered/nested observations (i.e., dependent observations) on the results and interpretation of a data analysis. |
| Type of analysis | Predictive. |
| Research question | What is the probability that the TSV vote for females at 25°C with a clothing insulation value of 0.6 clo in a typical (i.e., average) building is lower than 0.5 (‘neutral’)? |
| Framework | Bayesian. |
| Assumptions | Random sample (multistage sampling): everyone in the population has an equal chance of being selected into the sample. |
| Limited random variability: large sample size. | |
| No confounding: the DAG includes all shared causes among the variables. | |
| No model error: perfect functional form specification. | |
| No measurement error: all variables are measured perfectly. | |
| Variables | Indoor air temperature (T): continuous variable [unit: °C] |
| Thermal resistance of clothing (C): continuous variable [unit: clo] | |
| Sex (S): categorical variable [‘1’ male; ‘2’ female] | |
| Building ID (B_id): index variable | |
| TSV (V): continuous but bounded variable with interval (0, 1) [‘0’ cold; ‘1’ hot] |
4.3.3 Results
The results of two fitted statistical models (defined above) are presented here.
brms syntax for the formula argument
In brms, the intercept can be specified by the syntax y ~ x or y ~ 1 + x in the formula argument. When doing so the corresponding prior is applied under the presupposition all the regressors (x) are mean centred. However, if the regressors are not mean centred (like in our case), the syntax ... ~ 0 + Intercept + ... should be used. With this syntax, the prior can be defined on the real intercept, directly. More information can be found in the brms reference manual.
#Fit the beta regression model (Model 1)
Model.1 <-
brm(data = sample.random,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T + C,
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T),
prior(exponential(1), class = phi)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q9.fit_Model1') #save model fit
#View of the model summary
summary(Model.1) Family: beta
Links: mu = logit; phi = identity
Formula: V ~ 0 + Intercept + S + T + C
Data: sample.random (Number of observations: 10000)
Draws: 4 chains, each with iter = 5000; warmup = 1000; thin = 1;
total post-warmup draws = 16000
Regression Coefficients:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept -7.24 0.14 -7.52 -6.95 1.00 5160 7398
S2 -0.12 0.02 -0.15 -0.08 1.00 9304 9550
T 0.31 0.00 0.30 0.32 1.00 5961 8738
C 0.88 0.10 0.68 1.08 1.00 6108 8966
Further Distributional Parameters:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
phi 5.23 0.07 5.10 5.37 1.00 8574 9239
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).#Fit the beta regression model (Model 2)
Model.2 <-
brm(data = sample.random,
family = Beta(link = 'logit'),
formula = V ~ 0 + Intercept + S + T + C + (1 | B_id),
#Define priors for specific parameters or classes of parameters
prior = c(prior(normal(0, 5), class = b, coef = Intercept),
prior(normal(0, 1), class = b, coef = S2),
prior(normal(0, 1), class = b, coef = C),
prior(normal(0, 1), class = b, coef = T),
prior(exponential(1), class = phi),
prior(normal(0, 1), class = sd)),
iter = 5000, warmup = 1000,
chains = 4, cores = 4,
seed = 2023, #for reproducibility
file = 'fit/Q9.fit_Model2') #save model fit
#View of the model summary
summary(Model.2) Family: beta
Links: mu = logit; phi = identity
Formula: V ~ 0 + Intercept + S + T + C + (1 | B_id)
Data: sample.random (Number of observations: 10000)
Draws: 4 chains, each with iter = 5000; warmup = 1000; thin = 1;
total post-warmup draws = 16000
Multilevel Hyperparameters:
~B_id (Number of levels: 500)
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
sd(Intercept) 0.78 0.03 0.73 0.83 1.00 1286 3047
Regression Coefficients:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept -8.29 0.41 -9.10 -7.50 1.00 665 1575
S2 -0.15 0.01 -0.17 -0.13 1.00 32155 11712
T 0.35 0.02 0.32 0.39 1.00 691 1491
C 0.96 0.06 0.83 1.08 1.00 31834 12409
Further Distributional Parameters:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
phi 17.97 0.26 17.46 18.49 1.00 35041 10764
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).The two models can be compared by approximate leave-one-out cross-validation (LOOCV) (Vehtari et al., 2017) where smaller values indicate a better fit. In particular, the LOO information criterion (LOOIC) for the two models and their difference is calculated and summarised in Table 4.2.
Code
#Calculate LOO criterion for Model 1
Model.1 <- add_criterion(Model.1, criterion = 'loo')
#Calculate LOO criterion for Model 2
Model.2 <- add_criterion(Model.2, criterion = 'loo')
#Compare the LOOIC for the two models
LOOIC <-
rbind(Model.2 = loo(Model.2)$estimates[3, ], #extract LOOIC and SE for Model 2
Model.1 = loo(Model.1)$estimates[3, ]) |> #extract LOOIC and SE for Model 1
data.frame() |>
rename(c(LOOIC = Estimate, se = SE)) |>
round(digits = 2)
#Compare the difference of the LOOIC for the two models
LOO <- loo_compare(Model.2, Model.1, criterion = 'loo') #compare the elpd_loo
LOOIC.diff <-
cbind(LOOIC_diff = LOO[, 1] * (-2), #calculate the LOOIC difference
se_diff = LOO[, 2] * 2) |> #calculate the SE difference
round(digits = 2)| Model | LOOIC | SE | LOOIC.diff1 | SE.diff2 |
|---|---|---|---|---|
| Model 2: GLMM model | -17680.26 | 137.04 | 0.00 | 0.00 |
| Model 1: GLM model | -6067.95 | 110.81 | 11612.31 | 157.58 |
| 1 LOOIC.diff is the difference between the two LOOIC scores. | ||||
| 2 SE.diff is the standard error of the LOOIC.diff. | ||||
The table shows that Model.2 has a significantly better fit (smaller LOOIC value) than Model.1 since the difference in LOOIC (i.e., LOOIC.diff) is very large (73.69 times the corresponding standard error, SE.diff).
Using the function predicted_draws(), we can calculate the predicted posterior distribution of the TSVs for Model.1 and Model.2 given the regressors of interest (i.e., females at 25°C with clothing insulation of 0.6 clo).
#Create a data frame with defined regressors
grid.pred <- expand.grid(S = '2', #female
T = 25,
C = 0.6)
#Calculate prediction Model 1
set.seed(2023) #set random number for reproducibility
Model.1_pred <-
Model.1 |>
predicted_draws(newdata = grid.pred)
#Calculate prediction Model 2
set.seed(2023) #set random number for reproducibility
Model.2_pred <-
Model.2 |>
predicted_draws(newdata = grid.pred,
re_formula = NA) #include no group-level effectsThe predicted posterior distribution of the TSVs for Model.2 (i.e., Model.2_pred) represent the predicted TSVs for an average building. These have been calculated by holding the group-level residual (i.e., the group random effect) at its mean of zero, that is, by setting re_formula = NA in the function predicted_draws(). On the other hand, the predicted posterior distribution of the TSVs for Model.1 (i.e., Model.1_pred) do not represent the predicted TSVs for an average building because the model does not model the variability among buildings. Consequently, with this model is not possible to answer the question of interest. These predicted probabilities could be thought of, to a certain extent, as that of females at 25°C with clothing insulation of 0.6 clo across buildings on average. However, if these probabilities are of interest, they should be calculated using Model.2. This can be done by adding the term B_id = unique(sample.random$B_id) in grid.pred and by setting re_formula = NULL in the function predicted_draws().
The results of the two models are then plotted in Fig. 4.2.
Code
#Custom function to capitalise first letter of a word
foo = function(x){
paste(toupper(substring(x, 1, 1)),
tolower(substring(x, 2, nchar(x))),
sep = '')
}
#Combine the predictions of the two models in one data frame
plot_preds <-
bind_rows(
'Model.2_pred' = Model.2_pred,
'Model.1_pred' = Model.1_pred,
.id = 'draw_type') |>
mutate(draw_type = fct_inorder(draw_type))
#Calculate the highest predictive density (HPD) for Model 1 and Model 2
mean_preds <-
plot_preds |>
group_by(draw_type, S) |>
mean_hdi(.prediction) |>
mutate(.interval = 'hpd')
#Calculate the probability (percentage of area) that the predicted TSVs are lower and higher than 0.5 (i.e., neutral TSV)
perc.area <-
rbind(data.frame(lower = mean(Model.1_pred$.prediction < 0.5),
higher = mean(Model.1_pred$.prediction > 0.5),
draw_type = 'Model.1_pred'),
data.frame(lower = mean(Model.2_pred$.prediction < 0.5),
higher = mean(Model.2_pred$.prediction > 0.5),
draw_type = 'Model.2_pred'))
#Plot
ggplot(plot_preds, aes(x = .prediction, fill = draw_type)) +
stat_halfeye(point_interval = mean_hdi,
.width = .95,
shape = 16,
point_size = 2.3,
point_color = 'black', point_fill = 'black',
color = 'black',
normalize = 'all',
alpha = 0.75) +
geom_text(data = mean_preds, mapping = aes(x = .prediction, y = 0.95, label = paste(foo(.point), '=', sprintf('%.2f',round(.prediction, digits = 2)))),
size = 9/.pt, hjust = 0.5) +
geom_text(data = mean_preds, mapping = aes(x = .prediction, y = .25, label = paste(sprintf('%.0f%%',.width*100), toupper(.interval))),
size = 9/.pt, hjust = 0.5) +
geom_text(data = mean_preds, mapping = aes(x = .lower, y = 0.10, label = paste(sprintf('%.2f', .lower))),
size = 9/.pt, hjust = 0.5) +
geom_text(data = mean_preds, mapping = aes(x = .upper, y = 0.10, label = paste(sprintf('%.2f', .upper))),
size = 9/.pt, hjust = 0.5) +
geom_vline(xintercept = 0.5, linetype = 'dashed', linewidth = 0.25, alpha = 0.5) +
geom_text(data = perc.area, mapping = aes(x = 0.55, y = 0.60, label = paste(sprintf('%.1f%%', lower*100), '<', 0.50, '<', sprintf('%.1f%%', higher*100))),
size = 9/.pt, hjust = 0.5) +
scale_x_continuous(breaks = seq(0, 1, length.out = 7),
labels = c('Cold\n(0)', '', '', 'Neutral\n(0.5)', '', '', 'Hot\n(1)'),
limits = c(0, 1)) +
scale_fill_manual('Models', values = c('#F65314', '#7CBB00'),
breaks = c('Model.1_pred', 'Model.2_pred'),
labels = c('Model 1 (GLM)', 'Model 2 (GLMM)')) +
labs(x = expression('Predicted TSV'),
y = expression('Probability density')) +
facet_grid(. ~ factor(draw_type, levels=c('Model.1_pred','Model.2_pred'), labels = c('Model 1 (GLM)', 'Model 2 (GLMM)'))) +
theme(legend.position = 'none',
axis.ticks = element_line(linewidth = 0.25),
axis.title.x = element_text(margin = unit(c(2, 0, 0, 0), 'mm')),
axis.title.y = element_text(margin = unit(c(0, 2, 0, 0), 'mm'))
)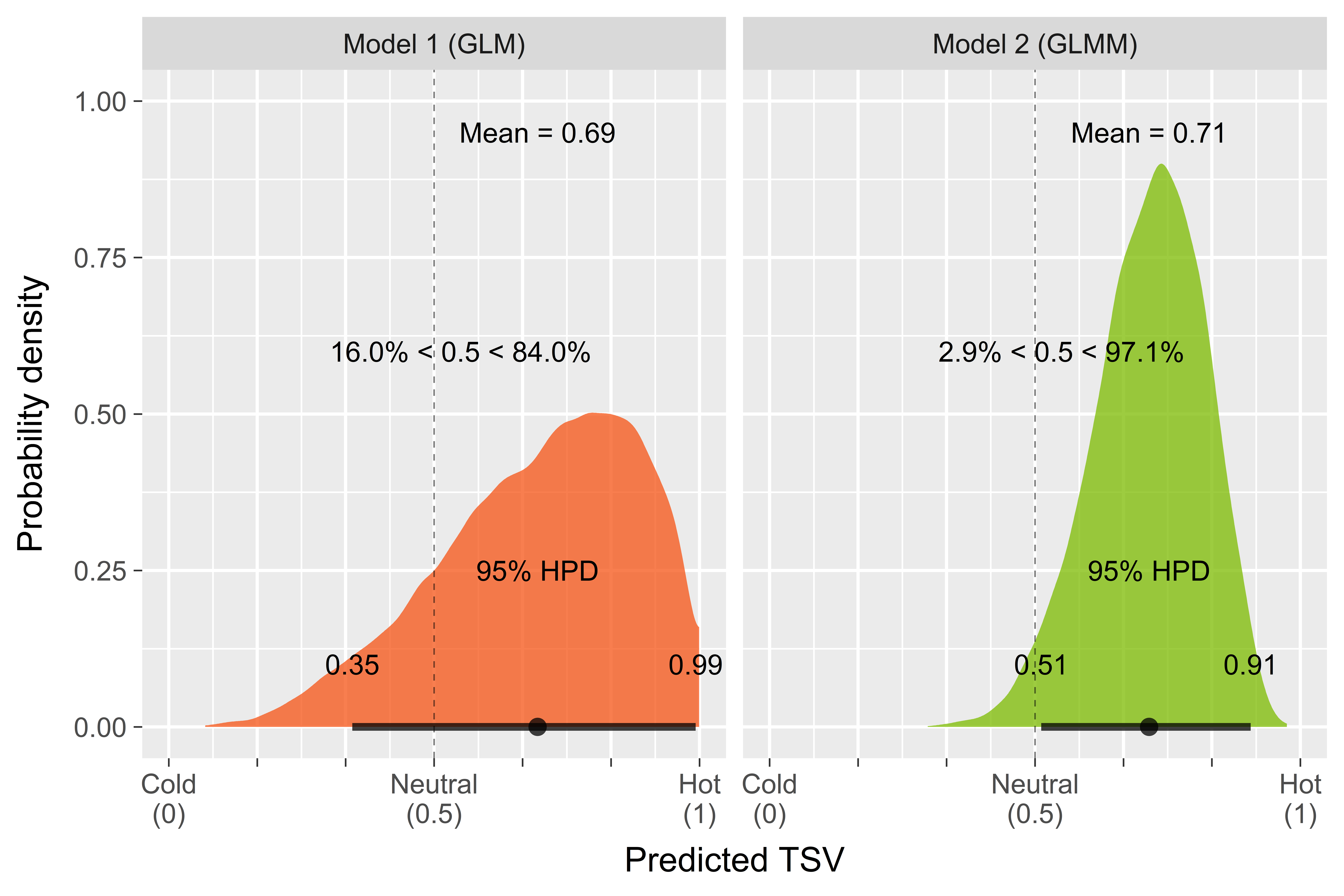
Fig. 4.2 shows the predicted posterior distribution of the TSVs for Model.1 (orange) and Model.2 (green), respectively. The black line and dot at the bottom of each distribution represent the highest predictive density (HPD) and the mean, respectively. These intervals (i.e., the black lines) are defined here to span over 95% of the distribution and represent the 95% HPD. Using Model.1, the probability that the thermal sensation vote for females at 25°C with a clothing insulation value of 0.6 clo is lower than 0.5 (‘neutral’) is 16.0%. However, this is not the probability for a typical (i.e., average) building, leading to the wrong answer to our question of interest. Model.2, on the other hand, can calculate the predicted TSVs for an average building. From this model, the probability that the thermal sensation vote for females at 25°C with a clothing insulation value of 0.6 clo in a typical (i.e., average) building is lower than 0.5 (‘neutral’) is 2.9%.
Session info
Version information about R, the OS and attached or loaded packages.
R version 4.2.3 (2023-03-15 ucrt)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 22631)
Matrix products: default
locale:
[1] LC_COLLATE=English_United Kingdom.utf8
[2] LC_CTYPE=English_United Kingdom.utf8
[3] LC_MONETARY=English_United Kingdom.utf8
[4] LC_NUMERIC=C
[5] LC_TIME=English_United Kingdom.utf8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] EnvStats_2.8.1 brms_2.21.0 Rcpp_1.0.12 tidybayes_3.0.6
[5] lubridate_1.9.3 forcats_1.0.0 stringr_1.5.1 dplyr_1.1.4
[9] purrr_1.0.2 readr_2.1.5 tidyr_1.3.1 tibble_3.2.1
[13] tidyverse_2.0.0 gt_0.11.0 data.table_1.15.4 ggplot2_3.5.1
[17] dagitty_0.3-4 ggdag_0.2.12
loaded via a namespace (and not attached):
[1] colorspace_2.1-0 estimability_1.5.1 QuickJSR_1.3.1
[4] rstudioapi_0.16.0 farver_2.1.2 rstan_2.32.6
[7] graphlayouts_1.1.1 ggrepel_0.9.5 svUnit_1.0.6
[10] fansi_1.0.6 mvtnorm_1.2-4 xml2_1.3.6
[13] bridgesampling_1.1-2 codetools_0.2-19 cachem_1.1.0
[16] knitr_1.48 polyclip_1.10-6 bayesplot_1.11.1
[19] jsonlite_1.8.8 ggdist_3.3.2 ggforce_0.4.2
[22] compiler_4.2.3 emmeans_1.10.3 backports_1.4.1
[25] Matrix_1.6-5 fastmap_1.2.0 cli_3.6.2
[28] tweenr_2.0.3 htmltools_0.5.8.1 tools_4.2.3
[31] igraph_2.0.3 coda_0.19-4.1 gtable_0.3.5
[34] glue_1.7.0 reshape2_1.4.4 posterior_1.6.0
[37] V8_4.4.2 vctrs_0.6.5 nlme_3.1-162
[40] ggraph_2.2.1 tensorA_0.36.2.1 xfun_0.46
[43] timechange_0.3.0 lifecycle_1.0.4 MASS_7.3-58.2
[46] scales_1.3.0 tidygraph_1.3.1 hms_1.1.3
[49] Brobdingnag_1.2-9 parallel_4.2.3 inline_0.3.19
[52] yaml_2.3.9 curl_5.2.1 memoise_2.0.1
[55] gridExtra_2.3 loo_2.8.0 StanHeaders_2.32.10
[58] sass_0.4.9 stringi_1.8.4 checkmate_2.3.1
[61] boot_1.3-28.1 pkgbuild_1.4.4 rlang_1.1.4
[64] pkgconfig_2.0.3 matrixStats_1.3.0 distributional_0.4.0
[67] evaluate_0.24.0 lattice_0.20-45 rstantools_2.4.0
[70] htmlwidgets_1.6.4 labeling_0.4.3 tidyselect_1.2.1
[73] plyr_1.8.9 magrittr_2.0.3 R6_2.5.1
[76] generics_0.1.3 pillar_1.9.0 withr_3.0.0
[79] abind_1.4-5 arrayhelpers_1.1-0 utf8_1.2.4
[82] tzdb_0.4.0 rmarkdown_2.27 viridis_0.6.5
[85] grid_4.2.3 digest_0.6.36 xtable_1.8-4
[88] RcppParallel_5.1.8 stats4_4.2.3 munsell_0.5.1
[91] viridisLite_0.4.2 We have selected only a few variables of interest to limit the complexity of the DAG and shift the focus to statistical data analysis.↩︎
The scientific model is generally an approximation (a best guess) of the unknown data-generating process. As such, it would have some inaccuracies/errors, which will inevitably affect the data analysis.↩︎
In all simulation examples, we relied on the regression framework (ordinal or beta regression) within the frequentist or Bayesian approach. However, other approaches are available and could be used depending on the question and the data. It is the researcher’s responsibility to justify the choice of the statistical framework (and any decisions made during the modelling process) to answer the specific question of interest.↩︎