3 Simulation example Q.8
In this simulation example, the highlighted data analysis pitfall is variable selection. Specifically, it shows the implication of variable selection on the result of the data analysis.
3.1 Data-generating process
The data-generating process is the specific process that generates the observed data. In reality, this process is unknown and understanding it is, in a general sense, the ultimate aim of science. Here, we assume this process is known and describe it via a directed acyclic graph (DAG) in Fig. 3.1. In this DAG,1 indoor air temperature \((\text{T})\) and biological sex \((\text{S})\) influence thermal sensation vote \((\text{V})\) both directly and indirectly, passing through clothing insulation \((\text{C})\) In addition, thermal sensation vote \((\text{V})\) is influenced indirectly by outdoor air temperature \((\text{O})\) through indoor air temperature \((\text{T})\) and clothing insulation \((\text{C})\).
Code
dag_coords <-
data.frame(name = c('T', 'C', 'S', 'V', 'O'),
x = c(1, 2, 3, 2, 1.5),
y = c(1.2, 1.2, 1.2, 1, 1.3))
DAG.08 <-
dagify(C ~ S,
C ~ T,
C ~ O,
T ~ O,
V ~ S + C + T,
coords = dag_coords)
ggplot(data = DAG.08, aes(x = x, y = y, xend = xend, yend = yend)) +
geom_dag_text(colour='black', size = 8, parse = TRUE,
family = c('mono'),
label = c(expression(bold(C)), expression(bold(O)), expression(bold(S)), expression(bold(T)), expression(bold(V)))) +
geom_dag_edges(arrow_directed = grid::arrow(length = grid::unit(8, 'pt'), type = 'open'),
family = c('mono'),
fontface = c('bold')) +
coord_cartesian(xlim = c(1, 3), ylim = c(1, 1.3)) +
theme_dag()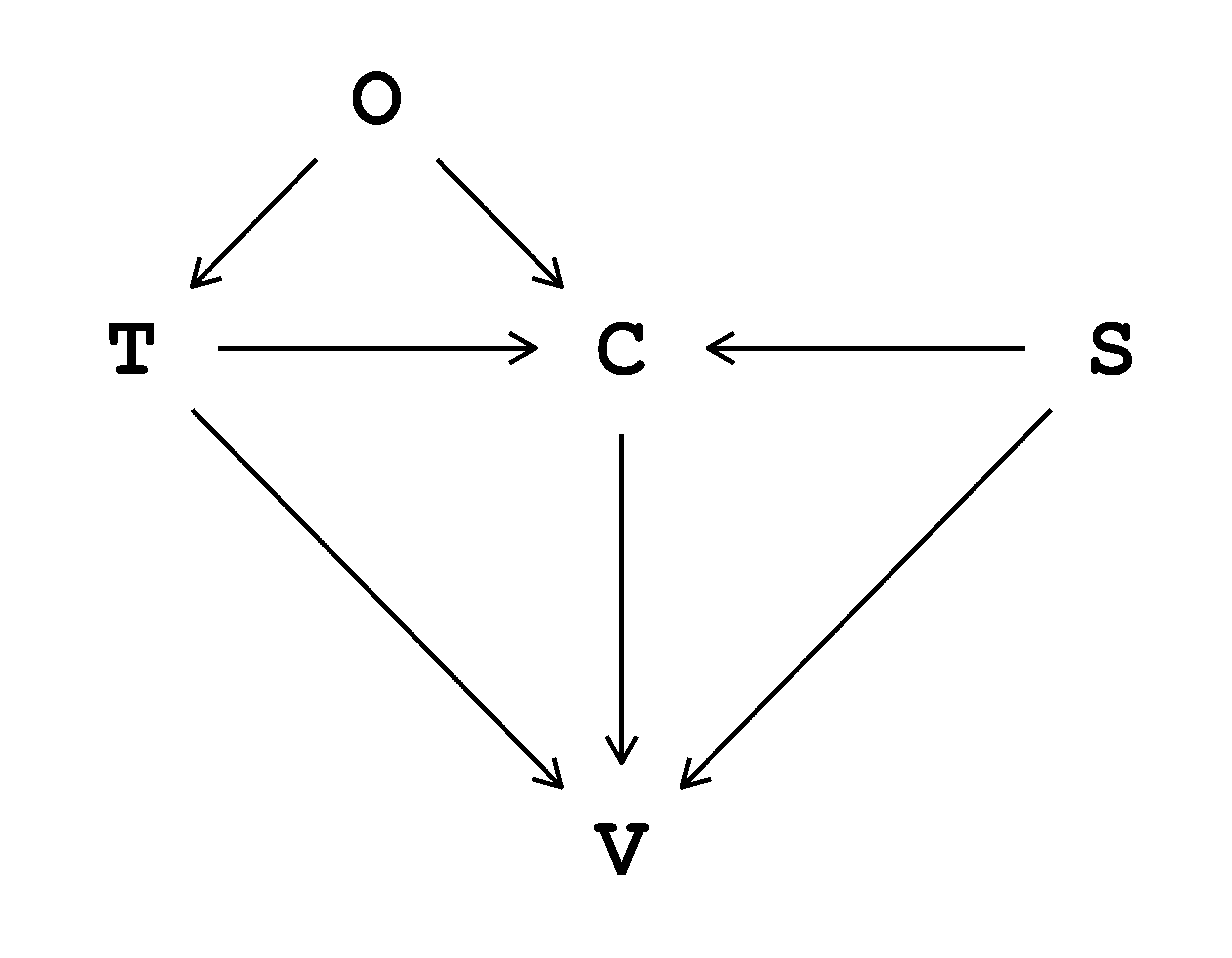
The DAG in Fig. 3.1 can be written as:
- \(T \sim f_{T}(O)\), read as ‘indoor air temperature \((\text{T})\) is some function of outdoor air temperature \((\text{O})\)’.
- \(C \sim f_{C}(S, T, O)\), read as ‘clothing insulation \((\text{C})\) is some function of biological sex \((\text{S})\), indoor air temperature \((\text{T})\) and outdoor air temperature \((\text{O})\)’.
- \(V \sim f_{V}(S, T, C)\), read as ‘thermal sensation vote \((\text{V})\) is some function of biological sex \((\text{S})\), indoor air temperature \((\text{T})\) and clothing insulation \((\text{C})\)’.
3.2 Synthetic data set
In a DAG, an omitted arrow between two variables signifies the absence of direct influence of one variable on another. In contrast, a present arrow does not provide any information about the magnitude of the effect or its functional form. As such, all relationships are qualitative. However, to generate synthetic data, we need to quantitatively define the relationship between the variables. These relationships are defined within the custom function data.sim_Q08.beta(). This function takes as input the sample size N and generates synthetic data according to the DAG in Fig. 3.1.
data.sim_Q08.beta <- function(N) {
#Simulate outdoor air temperature
O <- rbeta(N, shape1=2, shape2=2) * (30-(5)) + (5)
#Simulate indoor air temperature
T <- rnorm(N, mean = 19.87 + 7.17*inv_logit(-4.88 + 0.27*O), sd = 1)
#Simulate biological sex
S <- factor(sample(1:2, size=N, replace=TRUE)) #S = 1 'male'; S = 2 'female'
#Simulate clothing insulation
C <- rlnormTrunc(N, meanlog = ifelse(S=='1', 0.25 + 0.00 -0.01*T -0.02*O, 0.25 + 0.03 -0.01*T -0.02*O), sdlog = 0.15, min = 0.3 , max = 2)
#Simulate thermal sensation vote
V <- sim_TSV.beta(n=N, sex=S, temp=T, clo=C, b_int=-8.65, b_sex=c(0,-0.16), b_temp=0.37, b_clo=0.88, phi=17.9)
return(data.table(S, O, T, C, V)) #return a data.table with the simulated values
}Specifically:
Outdoor air temperature \((\text{O})\) is randomly sampled from a beta distribution with the two shape parameters \(\alpha\) (i.e.,
shape1 = 2) and \(\beta\) (i.e.,shape2 = 2) set to 2. Since the values obtained are defined only on the interval \((0,1)\), they are rescaled to be within 5°C and 30°C.Indoor air temperature \((\text{T})\) is randomly sampled from a normal distribution. The mean is a non-linear of function of outdoor air temperature (i.e.,
mean = 19.87 + 7.17*inv_logit(-4.88 + 0.27*O)) and the standard deviation is set to 1°C (i.e.,sd = 1). More information on the custom functioninv_logit()can be found in Appendix A.3.Biological sex \((\text{S})\) is randomly sampled with replacement from a vector of factors
1(i.e., male) and2(i.e., female).Clothing insulation \((\text{C})\) is randomly sampled from a truncated lognormal distribution. The mean is on the log scale (i.e.,
meanlog) and is a linear of function of biological sex \((\text{S})\) and indoor air temperature \((\text{T})\): for male (i.e.,S = 1) ismeanlog = 0.25 + 0.00 -0.01*T -0.02*Owhile for female (i.e.,S = 2) ismeanlog = 0.25 + 0.03 -0.01*T -0.02*O. The standard deviation is on the log scale and is a constant, fixed at 0.15 (i.e.,sdlog = 0.15). The parametersminandmaxare the assumed lower and upper bounds for clothing, fixed to 0.3 clo (i.e.,min = 0.3) and 2 clo (i.e.,max = 2), respectively.Thermal sensation votes \((\text{V})\) are generated from the custom function
sim_TSV.beta(), which is a function of biological sex (sex), indoor air temperature (temp), and clothing insulation (clo). The termb_intis the coefficient for the intercept (set to-8.65),b_sexis the coefficient for biological sex (set to0for males and-0.16for females),b_tempis the coefficient for indoor air temperature (set to0.37),b_clois the coefficient for clothing insulation (set to0.88), andphiis the precision parameter (set to17.9). More information on the custom functionsim_TSV.beta()can be found in Appendix A.3.
From this data generation mechanism, we simulated the target population, which consists of one million observations.
set.seed(2023) #set random number for reproducibility
#Simulate the population
Q08_population <- data.sim_Q08.beta(N = 1e6)
#View the data frame
Q08_population S O T C V
<fctr> <num> <num> <num> <num>
1: 1 16.909373 23.77877 0.6653989 0.4797464
2: 1 10.976568 21.30679 0.7348235 0.7770725
3: 1 6.988682 20.20013 0.6034874 0.3767384
4: 1 16.189995 22.53826 0.9155064 0.5659627
5: 2 13.141573 22.25360 0.8107148 0.5447163
---
999996: 2 19.393655 23.71747 0.5423450 0.6987992
999997: 2 8.744840 20.77339 1.0360876 0.6758423
999998: 2 10.776173 21.00722 0.6345823 0.3928896
999999: 1 14.702045 19.75479 1.0808817 0.3429427
1000000: 2 10.608257 21.63439 0.9771088 0.6102091From this population, we obtained a data set of ten thousand observations using simple random sampling.
set.seed(2023) #set random number for reproducibility
#Sample one data set of 10,000 observations
sample.random <-
Q08_population |>
slice_sample(n = 1e4) #take a simple random sample of size 10,000
#View the data frame
sample.random S O T C V
<fctr> <num> <num> <num> <num>
1: 1 9.04218 19.91598 0.7814319 0.2895530
2: 1 17.55348 23.22257 0.7151752 0.7137601
3: 2 18.64362 23.58910 0.8566554 0.7190638
4: 1 11.39534 21.87152 0.9136508 0.5784983
5: 2 11.65194 20.32116 0.9284893 0.3499993
---
9996: 2 17.74506 24.63568 0.8569835 0.7868010
9997: 2 13.31223 20.01855 0.9485145 0.4642949
9998: 1 14.71099 21.29476 0.8553684 0.4870056
9999: 1 27.94277 26.20721 0.5505871 0.9160256
10000: 2 15.08557 22.87278 0.8130227 0.5238597A summary of descriptive statistics for all variables in the data sets is given below. Detailed information can be found in Q.8 Summary.
summary(sample.random) S O T C V
1:4919 Min. : 5.164 Min. :16.46 Min. :0.3553 Min. :0.03532
2:5081 1st Qu.:13.138 1st Qu.:21.43 1st Qu.:0.6364 1st Qu.:0.46584
Median :17.472 Median :23.15 Median :0.7297 Median :0.62743
Mean :17.459 Mean :23.24 Mean :0.7443 Mean :0.61175
3rd Qu.:21.752 3rd Qu.:25.02 3rd Qu.:0.8369 3rd Qu.:0.76999
Max. :29.843 Max. :29.43 Max. :1.4477 Max. :0.99833 3.3 Data analysis
In a scientific context, data analysis should be performed to model the phenomenon under scrutiny and not fitting curves to data. The data are a means of obtaining information about that phenomenon to answer the specific question of interest. In this example, we assumed that the objective of the data analysis is to answer the following causal question: ‘What is the total causal effect of decreasing the indoor air temperature from 22°C to 20°C on the thermal sensation vote?’
3.3.1 Scientific model
The first step to address a scientific question is to define the scientific model. A scientific model is essentially a representation of a specific phenomenon that is being studied. To achieve this, we defined a DAG. A DAG generally consists of letter-and-arrow pictures summarising the researcher’s scientific knowledge about the phenomenon under study. Since we know the data-generating process (we defined it), the scientific model equates to the data-generating process outlined in Fig. 3.1. In reality, however, it would not.2
3.3.2 Statistical model
A statistical model is a mathematical representation of the scientific model —which in turn represents the data-generating process— that aims to answer the particular question of interest. To address the question, the synthetic data sets are analysed using beta regression within a frequentist framework.3
To estimate the effect of interest, we need to select the variables to adjust for: the adjustment set. This selection is done in two ways: using causal thinking and backward selection with the AIC criterion. Using causal thinking (assuming that the structure of the scientific model is correct), we can use the adjustmentSets() function to identify the adjustment set that (asymptotically) allow the unbiased estimation of the effect of interest.
adjustmentSets(DAG.08,
exposure = 'T',
outcome = 'V',
type = 'all',
effect = 'total',
max.results = Inf){ O }
{ O, S }There are two possible adjustment sets to estimate the total causal effect of indoor air temperature: include outdoor air temperature \((\text{O})\) (Model 1) or include outdoor air temperature \((\text{O})\) and sex \((\text{S})\) (Model 2). The statistical model for ‘Model 1’ can then be written as:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Beta}(\mu_{i}, \phi)\\ \operatorname{logit}(\mu_{i}) &= \eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{T} \mathrm{T}_{i} + \beta_{O} \mathrm{O}_{i} \\ \end{aligned} \tag{3.1}\]
whereas the statistical model for ‘Model 2’ can then be written as:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Beta}(\mu_{i}, \phi)\\ \operatorname{logit}(\mu_{i}) &= \eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{S} (\mathrm{S}_i) + \beta_{T} \mathrm{T}_{i} + \beta_{O} \mathrm{O}_{i} \\ \end{aligned} \tag{3.2}\]
To perform backward selection with the AIC criterion, the initial statistical model include all four regressors, leading to the following full model:
\[ \begin{aligned} \mathrm{V}_{i} &\sim \operatorname{Beta}(\mu_{i}, \phi)\\ \operatorname{logit}(\mu_{i}) &= \eta_{i}\\ \eta_{i} &= \alpha_{0} + \beta_{S} (\mathrm{S}_i) + \beta_{T} \mathrm{T}_{i} + \beta_{C} \mathrm{C}_{i} + \beta_{O} \mathrm{O}_{i} \\ \end{aligned} \tag{3.3}\]
In Eq. 5.1, Eq. 5.2 and Eq. 3.3, the conditional distribution of the response variable \(\mathrm{Y}_{i}\) for the \(i^{th}\) observation is assumed to follow a beta distribution with mean \(\mu_{i}\) and precision parameter \(\phi\). The parameter \(\eta_{i}\) is the linear predictor, where \(\alpha_{0}\) is the intercept term and \(\beta_{S}\), \(\beta_{T}\), \(\beta_{C}\), \(\beta_{O}\) are the regression coefficients for biological sex \((\text{S})\), indoor air temperature \((\text{T})\), clothing insulation \((\text{C})\), and outdoor air temperature \((\text{O})\), respectively.
A summary description of the simulation example is provided in Table 3.1.
| Pitfall | Ignore the implications of variable selection on the results and interpretation of a data analysis. |
| Type of analysis | Causal. |
| Research question | What is the total causal effect of decreasing the indoor air temperature from 22°C to 20°C on the TSV? |
| Framework | Frequentist. |
| Assumptions | Random sample (simple random sampling): everyone in the population has an equal chance of being selected into the sample. |
| Limited random variability: large sample size. | |
| Independence of observations: each observation represents independent bits of information. | |
| No confounding: the DAG includes all shared causes among the variables. | |
| No model error: perfect functional form specification. | |
| No measurement error: all variables are measured perfectly. | |
| Variables | Outdoor air temperature (O): continuous variable [unit: °C] |
| Indoor air temperature (T): continuous variable [unit: °C] | |
| Thermal resistance of clothing (C): continuous variable [unit: clo] | |
| Sex (S): categorical variable [‘1’ male; ‘2’ female] | |
| TSV (V): continuous but bounded variable with interval (0, 1) [‘0’ cold; ‘1’ hot] |
3.3.3 Results
The results of three fitted statistical models (defined above) are presented here.
#Fit the beta regression model with T and O (Model 1)
Model.1 <-
glmmTMB(formula = V ~ 1 + T + O,
data = sample.random,
family = beta_family(link = "logit"),
dispformula = ~ 1)
#View of the model summary
summary(Model.1) Family: beta ( logit )
Formula: V ~ 1 + T + O
Data: sample.random
AIC BIC logLik deviance df.resid
-16706.3 -16677.5 8357.2 -16714.3 9996
Dispersion parameter for beta family (): 16.8
Conditional model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -7.741883 0.087330 -88.65 < 2e-16 ***
T 0.367373 0.004982 73.74 < 2e-16 ***
O -0.015928 0.001944 -8.19 2.51e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#Fit the beta regression model with S, T and O (Model 2)
Model.2 <-
glmmTMB(formula = V ~ 1 + S + T + O,
data = sample.random,
family = beta_family(link = "logit"),
dispformula = ~ 1)
#View of the model summary
summary(Model.2) Family: beta ( logit )
Formula: V ~ 1 + S + T + O
Data: sample.random
AIC BIC logLik deviance df.resid
-16920.0 -16883.9 8465.0 -16930.0 9995
Dispersion parameter for beta family (): 17.2
Conditional model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -7.664684 0.086602 -88.50 < 2e-16 ***
S2 -0.148962 0.010094 -14.76 < 2e-16 ***
T 0.367161 0.004933 74.43 < 2e-16 ***
O -0.015688 0.001925 -8.15 3.67e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#Fit the beta regression model with S, T, C and O (Model 3)
Model.3 <-
buildglmmTMB(formula = V ~ 1 + S + T + C + O,
data = sample.random,
buildmerControl = buildmerControl(direction = c('order','backward'),
crit = 'AIC', elim = 'AIC',
family = beta_family(link = "logit"),
quiet = TRUE))
#View of the model summary
summary(Model.3) Family: beta ( logit )
Formula: V ~ 1 + T + C + S
Data: sample.random
AIC BIC logLik deviance df.resid
-17352.1 -17316.1 8681.1 -17362.1 9995
Dispersion parameter for beta family (): 18
Conditional model:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -8.681568 0.088278 -98.34 <2e-16 ***
T 0.370266 0.002917 126.93 <2e-16 ***
C 0.919232 0.040816 22.52 <2e-16 ***
S2 -0.170927 0.009949 -17.18 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1The backward selection with the AIC criterion performed in Model 3 has selected the variables indoor air temperature \((\text{T})\), clothing insulation \((\text{C})\) and sex \((\text{S})\). However, this is the wrong adjustment set,4 and it will not allow us to estimate the total causal effect of indoor air temperature. The AIC criterion is a predictive criterion and, as such, is a tool for pure predictive tasks (i.e., prediction in the absence of intervention). Here, the question of interest is causal, that is, prediction in the presence of intervention (i.e., an answer to a ‘what if?’ question).
Table 3.2 compares the three models in terms of their AIC: the model selected by backward selection (Model 3) has the best AIC (i.e., the lowest value) and, as such, it is the best model for a pure predictive task. However, it will be the wrong model to answer the question of interest.
| Variable selection strategy | Selected model | df1 | AIC |
|---|---|---|---|
| Causal thinking | Model 1: V ~ T + O | 4 | -16706.3 |
| Model 2: V ~ S + T + O | 5 | -16920.0 | |
| Backward selection | Model 3: V ~ S + T + C | 5 | -17352.1 |
| 1 df stands for degrees of freedom. | |||
The choice is, therefore, between Model 1 and Model 2. The minimal adjustment set (i.e., the one that uses as few variables as possible) includes only outdoor air temperature \((\text{O})\). In this view, adjusting for (i.e., conditioning on) sex is optional. However, conditioning on sex can help detect the effect of indoor air temperature by explaining some residual variations in thermal sensation vote, thus improving precision. For this reason, we selected the adjustment set of Model 2.
To calculate the causal effect of interest, we will use the potential outcomes framework (Imbens and Rubin, 2015; Neyman, Jerzy et al., 1990; Rubin, 1974). Our outcome of interest is the TSV \((\text{V})\), which we can write as \(Y\). Since \(Y\) varies across \(i\) persons, we can denote each participant’s TSV as \(Y_i\). Our (well-defined) intervention is a change from 22°C to 20°C of the indoor air temperature. For notation purposes, we can let \(0\) stand for indoor temperature at 22°C and \(1\) stand for indoor temperature at 20°C. Consequently, we can write \(Y_i^0\) for the \(i\)-th person’s outcome if they were exposed to an indoor temperature of 22°C and \(Y_i^1\) for the \(i\)-th person’s outcome if they were exposed to an indoor temperature of 20°C. Therefore, we can define the causal effect \(\tau\) of the change in temperature for the \(i\)-th person as \(\tau_i=Y_i^1-Y_i^0\). The average treatment effect (ATE) can be computed as
\[ \begin{aligned} \tau_{ATE}=\mathbb{E}(\tau_i)=\mathbb{E}(Y_i^1-Y_i^0) \end{aligned} \tag{3.4}\]
where \(\mathbb{E}(~)\) is the expectation operator. This formula implies that the average treatment effect in the population equals the average of each person’s treatment effect.5 However, the quantity \(\tau_i=Y_i^1-Y_i^0\) is unobservable. In fact, for each person, we observe, at best, only one of the two terms necessary to calculate the individual causal effect, \(\tau_i\). We could observe \(Y_i^1\) or \(Y_i^0\) but not both. Therefore, we always miss at least half of the data. This is called the fundamental problem of causal inference (Holland, 1986).
A solution is to use our regression model (Model 2) to compute the counterfactual estimates \(\hat{Y}_i^1\) and \(\hat{Y}_i^0\), which we can then use to calculate the estimate for \(\mathbb{E}(Y_i^1-Y_i^0)\), that is, \(\hat{\tau}_{ATE}=\mathbb{E}(\hat{Y}_i^1-\hat{Y}_i^0)\). In our example, however, we also have the outdoor air temperature \((\text{O})\) and sex \((\text{S})\) that we have to consider. To do so, we need to expand the framework. Specifically, if we assume that \(\mathbf{W}_i\) is a vector of regressors (the values for which vary across the participants), we can write the ATE as
\[ \begin{aligned} \tau_{ATE}=\mathbb{E}(Y_i^1-Y_i^0\ |\ \mathbf{W}_i) \end{aligned} \tag{3.5}\]
This formula implies that the average treatment effect in the population equals the average of each person’s treatment effect, computed conditional on their values for the vector of regressors \(\mathbf{W}_i\). This method is called g-computation algorithm formula, which, in simple terms, can be described as follows:
- Use the observed data to fit the regression model (i.e.,
Model 2) with \(Y\) as regressand (i.e., \(\text{V}\)), \(X\) as treatment (i.e., \(\text{T}\)), and \(\mathbf{W}\) as the adjustment set (i.e., \(\text{O}\) and \(\text{S}\)). - Create a new data set identical to the original data, but where \(X = 0\) (i.e., \(\text{T}=22°\text{C}\)) in every row.
- Create a new data set identical to the original data, but where \(X = 1\) (i.e., \(\text{T}=20°\text{C}\)) in every row.
- Use the model from Step 1 (i.e.,
Model 2) to compute adjusted predictions in the two counterfactual data sets from Steps 2 and 3. - The quantity of interest is the difference between the means of adjusted predictions in the two counterfactual data sets, that is, \(\hat{\tau}_{ATE}=\mathbb{E}(\hat{Y}_i^1-\hat{Y}_i^0\ |\ \mathbf{W}_i)\).
It goes without saying that the veracity of this calculation’s result is contingent on the model and, consequently, all the assumptions the model relies upon.
Causal inference requires additional assumptions, generally called identifiability conditions. These conditions will vary depending on the framework used for causal inference. For example, for the potential outcome framework, these assumptions are:
- the ‘stable-unit-treatment-value assumption’ (SUTVA), which can be divided into two main components: ‘no interference’ and ‘consistency’;
- ‘positivity’ (also called ‘overlap’);
- ‘exchangeability’ (also known as ‘ignorability’ or ‘unconfoundedness’).
An in-depth discussion of causal inference is beyond the scope of this notebook. For more information, the reader is referred to the book by Hernán and Robins (Hernán and Robins, 2020).
Using the function predictions(), we can produce unit-level estimates (i.e., one specific prediction per observation) for the TSV \((\text{V})\). By default, the predictions() function makes one prediction per observation in the data set that was used to fit the model. However, we are only interested in the predictions of TSV at two specific levels of indoor air temperature (i.e., at 20°C and 22°C). To do so, we specified the variables = list(T = c(20, 22)) argument within the predictions() function. In this way the predictions() function calculate counterfactual predictions for \(\text{T}= 20\text{°C}\) and \(\text{T}= 22\text{°C}\) leaving outdoor air temperature \((\text{O})\) and sex \((\text{S})\) at their observed value. Therefore, the entire data set is duplicated twice: once for each of the variable values specified in variables (i.e., \(\text{T}= 20\text{°C}\) and \(\text{T}= 22\text{°C}\)).
#counterfactual variables
pred <-
predictions(Model.2,
variables = list(T = c(20, 22)), #list of names identify the subset of variables of interest and their values
vcov = TRUE, #variance-covariance matrix
re.form = NA, #used to silence a warning that is irrelevant to this model
type = 'response')
#View the result
pred
S O T Estimate Std. Error z Pr(>|z|) S 2.5 % 97.5 %
1 9.04 20 0.386 0.00244 158.5 <0.001 Inf 0.381 0.391
1 17.55 20 0.355 0.00396 89.6 <0.001 Inf 0.347 0.363
2 18.64 20 0.318 0.00411 77.3 <0.001 Inf 0.310 0.326
1 11.40 20 0.377 0.00241 156.4 <0.001 Inf 0.373 0.382
2 11.65 20 0.342 0.00233 147.1 <0.001 Inf 0.338 0.347
--- 19990 rows omitted. See ?avg_predictions and ?print.marginaleffects ---
2 17.75 22 0.496 0.00235 211.4 <0.001 Inf 0.492 0.501
2 13.31 22 0.514 0.00197 260.7 <0.001 Inf 0.510 0.518
1 14.71 22 0.545 0.00187 291.4 <0.001 Inf 0.542 0.549
1 27.94 22 0.494 0.00665 74.2 <0.001 Inf 0.481 0.507
2 15.09 22 0.507 0.00186 273.0 <0.001 Inf 0.503 0.510
Columns: rowid, rowidcf, estimate, std.error, statistic, p.value, s.value, conf.low, conf.high, V, S, O, T
Type: response As a result, pred contains 20,000 rows.
To compute the actual unit-level contrasts \(Y_i^1-Y_i^0\), we need to use the comparisons() function. By specifying the variables argument as before (i.e., variables = list(T = c(20, 22))), the comparisons() function calculate the counterfactual comparisons between the two levels of indoor air temperature (i.e., \(\text{T}= 20\text{°C}\) and \(\text{T}= 22\text{°C}\)) for each case in the original data set.
pred.contrast <-
comparisons(Model.2,
variables = list(T = c(20, 22)), #list of names identify the subset of variables of interest and their values
comparison = function(hi, lo) lo - hi,
vcov = TRUE, #variance-covariance matrix
re.form = NA, #used to silence a warning that is irrelevant to this model
type = 'response')
#View the result
pred.contrast
Term Contrast Estimate Std. Error z Pr(>|z|) S 2.5 % 97.5 %
T custom -0.181 0.00241 -75.1 <0.001 Inf -0.186 -0.176
T custom -0.179 0.00215 -83.4 <0.001 Inf -0.183 -0.175
T custom -0.175 0.00190 -92.0 <0.001 Inf -0.179 -0.171
T custom -0.181 0.00237 -76.2 <0.001 Inf -0.185 -0.176
T custom -0.178 0.00233 -76.4 <0.001 Inf -0.183 -0.173
--- 9990 rows omitted. See ?avg_comparisons and ?print.marginaleffects ---
T custom -0.175 0.00197 -89.1 <0.001 Inf -0.179 -0.171
T custom -0.177 0.00225 -79.0 <0.001 Inf -0.182 -0.173
T custom -0.180 0.00227 -79.2 <0.001 Inf -0.185 -0.176
T custom -0.175 0.00144 -121.4 <0.001 Inf -0.178 -0.172
T custom -0.177 0.00214 -82.4 <0.001 Inf -0.181 -0.172
Columns: rowid, term, contrast, estimate, std.error, statistic, p.value, s.value, conf.low, conf.high, predicted_lo, predicted_hi, predicted, V, S, T, O
Type: response However, as defined in Eq. 3.5, our estimand ATE is \(\mathbb{E}(Y_i^1-Y_i^0\ |\ \mathbf{W}_i)\). Therefore, we need to compute the average of those unit-level contrasts. Using the avg_comparisons() function, we can estimate the total average causal effect of decreasing the indoor air temperature from 22°C to 20°C.
pred.contrast_avg <-
avg_comparisons(Model.2,
variables = list(T = c(20, 22)), #list of names identify the subset of variables of interest and their values
comparison = function(hi, lo) lo - hi,
vcov = TRUE, #variance-covariance matrix
re.form = NA, #used to silence a warning that is irrelevant to this model
type = 'response')
#View the result
pred.contrast_avg
Term Contrast Estimate Std. Error z Pr(>|z|) S 2.5 % 97.5 %
T custom -0.177 0.002 -88.6 <0.001 Inf -0.181 -0.173
Columns: term, contrast, estimate, std.error, statistic, p.value, s.value, conf.low, conf.high
Type: response The estimate of the total average causal effect of decreasing the indoor air temperature from 22°C to 20°C is a reduction of -0.177 (95% CI [-0.181, -0.173]) of the mean thermal sensation vote. The critical aspect to remember is that all the variables added as the adjustment set are solely there to enable the estimation of the estimate of interest. Here, outdoor air temperature and sex are selected to yield the total causal effect of indoor air temperature, and nothing else. As such, these variables should not be interpreted. The interpretation of such variables is known as ‘Table 2 Fallacy’ (Westreich and Greenland, 2013).
Session info
Version information about R, the OS and attached or loaded packages.
R version 4.2.3 (2023-03-15 ucrt)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 22631)
Matrix products: default
locale:
[1] LC_COLLATE=English_United Kingdom.utf8
[2] LC_CTYPE=English_United Kingdom.utf8
[3] LC_MONETARY=English_United Kingdom.utf8
[4] LC_NUMERIC=C
[5] LC_TIME=English_United Kingdom.utf8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] EnvStats_2.8.1 marginaleffects_0.21.0 buildmer_2.11
[4] glmmTMB_1.1.9 tidybayes_3.0.6 lubridate_1.9.3
[7] forcats_1.0.0 stringr_1.5.1 dplyr_1.1.4
[10] purrr_1.0.2 readr_2.1.5 tidyr_1.3.1
[13] tibble_3.2.1 tidyverse_2.0.0 gt_0.11.0
[16] data.table_1.15.4 ggplot2_3.5.1 dagitty_0.3-4
[19] ggdag_0.2.12
loaded via a namespace (and not attached):
[1] nlme_3.1-162 insight_0.20.2 numDeriv_2016.8-1.1
[4] tensorA_0.36.2.1 tools_4.2.3 TMB_1.9.14
[7] backports_1.4.1 utf8_1.2.4 R6_2.5.1
[10] mgcv_1.8-42 colorspace_2.1-0 ggdist_3.3.2
[13] withr_3.0.0 gridExtra_2.3 tidyselect_1.2.1
[16] emmeans_1.10.3 curl_5.2.1 compiler_4.2.3
[19] cli_3.6.2 arrayhelpers_1.1-0 xml2_1.3.6
[22] sass_0.4.9 labeling_0.4.3 posterior_1.6.0
[25] scales_1.3.0 checkmate_2.3.1 mvtnorm_1.2-4
[28] digest_0.6.36 minqa_1.2.7 rmarkdown_2.27
[31] pkgconfig_2.0.3 htmltools_0.5.8.1 lme4_1.1-35.5
[34] fastmap_1.2.0 htmlwidgets_1.6.4 rlang_1.1.4
[37] rstudioapi_0.16.0 generics_0.1.3 farver_2.1.2
[40] svUnit_1.0.6 jsonlite_1.8.8 distributional_0.4.0
[43] magrittr_2.0.3 Matrix_1.6-5 Rcpp_1.0.12
[46] munsell_0.5.1 fansi_1.0.6 viridis_0.6.5
[49] abind_1.4-5 lifecycle_1.0.4 stringi_1.8.4
[52] yaml_2.3.9 ggraph_2.2.1 MASS_7.3-58.2
[55] grid_4.2.3 ggrepel_0.9.5 lattice_0.20-45
[58] graphlayouts_1.1.1 splines_4.2.3 hms_1.1.3
[61] knitr_1.48 pillar_1.9.0 igraph_2.0.3
[64] boot_1.3-28.1 estimability_1.5.1 glue_1.7.0
[67] evaluate_0.24.0 V8_4.4.2 vctrs_0.6.5
[70] nloptr_2.1.1 tzdb_0.4.0 tweenr_2.0.3
[73] gtable_0.3.5 polyclip_1.10-6 cachem_1.1.0
[76] xfun_0.46 ggforce_0.4.2 xtable_1.8-4
[79] tidygraph_1.3.1 coda_0.19-4.1 viridisLite_0.4.2
[82] memoise_2.0.1 timechange_0.3.0 We have selected only a few variables of interest to limit the complexity of the DAG and shift the focus to statistical data analysis.↩︎
The scientific model is generally an approximation (a best guess) of the unknown data-generating process. As such, it would have some inaccuracies/errors, which will inevitably affect the data analysis.↩︎
In all simulation examples, we relied on the regression framework (ordinal or beta regression) within the frequentist or Bayesian approach. However, other approaches are available and could be used depending on the question and the data. It is the researcher’s responsibility to justify the choice of the statistical framework (and any decisions made during the modelling process) to answer the specific question of interest.↩︎
Given the DAG, assuming air temperature and sex are conditioned on (i.e., included in the model), conditioning on clothing will statistically remove any association between indoor air temperature and thermal sensation vote influenced by clothing insulation. Stated analogously, it will block the indirect effect of indoor air temperature (i.e., the mediating path).↩︎
Here, we assumed that the calculated causal effect equals the population average treatment effect (PATE) because the sample is representative of the target population. However, if a sample does not refer to any population distribution of subject characteristics, the calculated quantity should be called sample average treatment effect (SATE), not PATE.↩︎